Cellular Genetics
Archive Page
This page is maintained as a historical record and is no longer being updated.
In 2025, the Cellular Genetics programme was renamed the Cellular Genomics programme.
To access the Sanger Institute’s current research in this is area, please visit the Cellular Genomics programme.
Overview
Mapping cells across time and space
Cellular Genetics scientists are key contributors to the Human Cell Atlas international consortium, which aims to create comprehensive reference maps of all human cells. To generate high-resolution, usable cell maps to benefit the global research community, we unite world-class wet- and dry- lab technologies and expertise under one roof at Sanger.
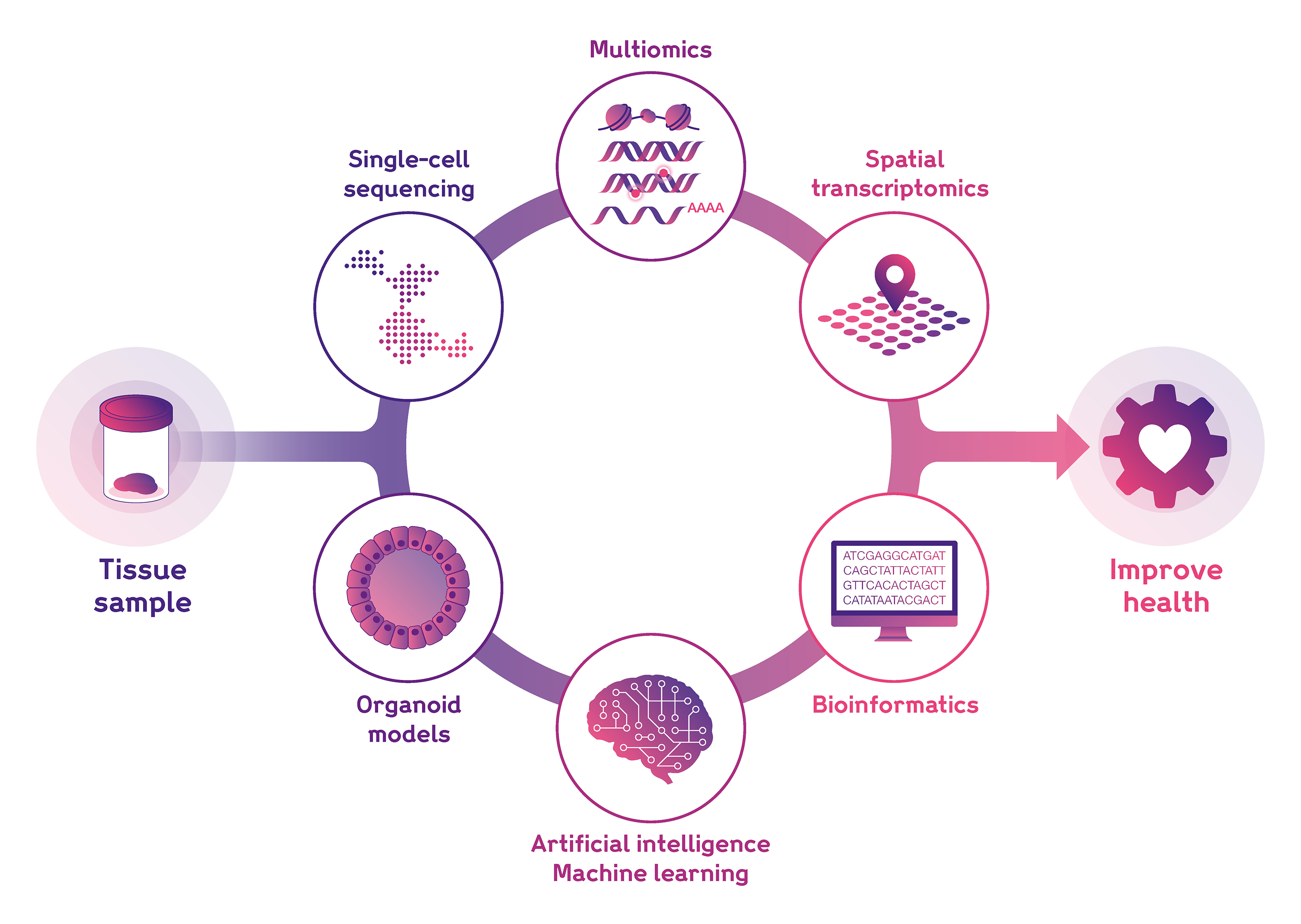
We utilise state-of-the-art methodologies to characterise multiple features of the cells and tissues comprising the human body, including:
- Single-cell transcriptomics
- Spatial transcriptomics
- Single-cell multiomics
- Bioinformatics and data science
- Artificial Intelligence and Machine Learning, including large language models and foundation models.
- In vitro models, including organoids
Our comprehensive cell maps allow us to delve into the intricate frameworks that govern cellular behaviour and interactions. By understanding the mechanisms by which cells assemble to form functioning tissues and organs, we uncover the design principles of the human body. Ultimately, our work aims to bridge the gap between genotype and phenotype, deciphering how the body interprets the genome to generate diverse cells and tissues.
With trillions of cells orchestrating precise interactions to maintain health, our research sheds light on the causes and consequences of disruption in disease, paving the way for transformative advances in medicine.
Our impact
Our pioneering research groups unlock new insights into disease mechanisms and therapeutic strategies by:
- Decoding the genetic and epigenetic mechanisms controlling human development and disease
- Investigating the cellular basis of immune disorders, for instance in the skin and kidneys
- Discovering the molecular drivers of childhood cancers to identify novel therapeutic targets
- Exploring the developmental origins and physiological functions of female reproductive tissues, and diseases such as endometriosis
- Analysing large-scale genomic datasets to drive discoveries in precision medicine and systems biology
- Developing wet-lab protocols and computational tools to accelerate scientific discoveries
Driving research culture and equity
We recognise that our single-cell genomics data is most valuable when easily accessible and usable by all. We are committed to democratising our data, ensuring that our scientific insights are available to the entire global research community.
We develop portals to make our datasets intuitive to browse and query for researchers, clinicians and educators globally. We also actively share our findings through pre-prints to ensure our discoveries and data can rapidly power further research, in line with our open science philosophy.
Wherever possible, we seek to study the full diversity of the global human population. We achieve this by ensuring representative samples in our research to promote data equity in both range and access. Furthermore, we actively engage with patient groups, ensuring that our research addresses their real-world needs.
Diversity and inclusivity are cornerstones of the Cellular Genetics Programme – not only in our research but in our researchers too. Our researchers bring a wide range of scientific expertise and lived experiences from around the world, enriching our science. We are proud that our inclusive research culture empowers our teams to drive forward transformative discoveries and innovative science.