Lotfollahi Group
Artificial Intelligence for Cell Engineering (AICE) Lab
Vision
Our vision is to create a digital version of every human cell in all organs, enabling an understanding of cell behaviour in the tissue and response to perturbations. We will seek to engineer resilient cells that resist diseases or reverse them by precisely modulating target cell behaviour to design personalised therapies.
This mission leverages state-of-the-art machine learning and artificial intelligence, high-throughput genomics and imaging, and perturbation experiments. Our team will be interdisciplinary, including computer scientists, bioinformaticians, and biologists to collaboratively drive our mission.
Biological systems
We will be working on experimental models to generate large-scale perturbation experiments to study:
- Engineering iPSC-cardiomyocytes that better match their in vivo counterparts.
- Modelling and modulating cellular interaction and response during neurulation. We study complex genetic interactions and their downstream gene regulatory networks leading to dysregulated inflammatory responses.
- We seek to understand which genetic variations and regulatory networks modulate T-cell activation and gene expression dynamics.
Understanding the system through Artificial Intelligence and High-Throughput Genomics
To create better cells, we must first understand their health and disease functions.
We will actively contribute to the Human Cell Atlas (HCA) project, aiming to build digital twins for each organ. Our approach will span multiple scales:
- starting with the behaviour of individual cells
- moving up to their interactions within tissues
- then their impact on overall organ function
- and, ultimately, the genetic variations that distinguish us at each level.
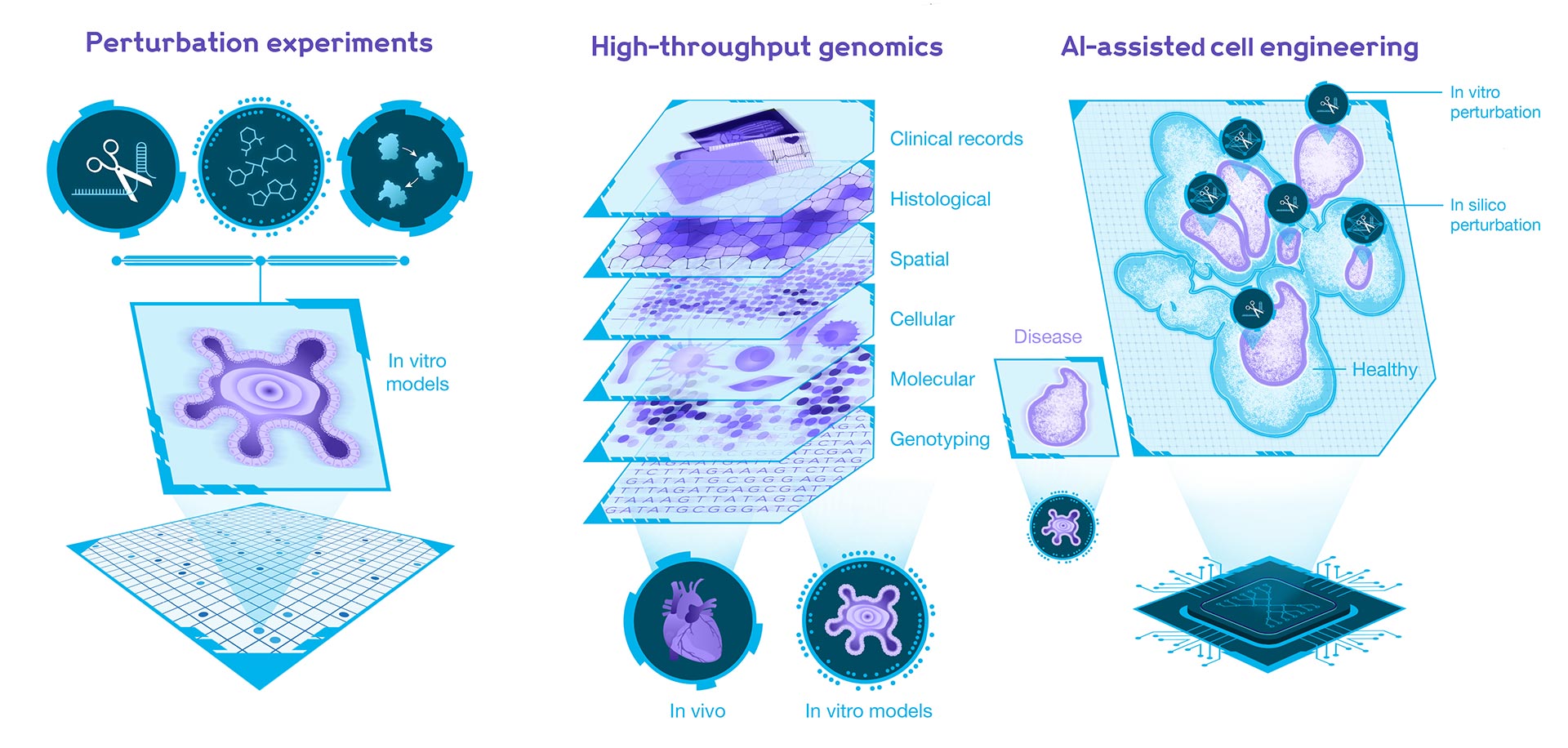
Modelling the system changes through Artificial Intelligence and Experimental Perturbations
Understanding a biological system necessitates understanding how individual components respond to perturbations, i.e., our cells.
The space of potential perturbations in biology is vast, ranging from genetic manipulation to small molecules. Observing the system’s behaviour in response to every possible perturbation is experimentally impractical. Therefore, we will combine machine learning with extensive perturbation experiments to identify the perturbations that induce the specific cellular behaviours of interest.
Leveraging AI, High-Throughput Genomics and Perturbations for Cell engineering and medical therapies
Once we understand normal and disease-related system behaviours and responses to perturbations, we can design enhanced cells and, potentially, whole organs.
We will overcome our in vitro systems’ accuracy limitations by integrating Human Cell Atlas data with our in-house data generation efforts. We will then apply Machine Learning to compare our in vitro systems with in vivo references to refine our cell engineering and organ generation methods.
This approach holds great potential for cell therapy, regenerative medicine, and personalised medicine.
