Cellular Genomics
Overview
Our strategy
A huge gap currently exists in our understanding of how genetic sequences instruct tissue differentiation during development, and how they influence patient presentation in the clinic. Tissue ecosystems are incredibly complex, comprising millions of cells of a remarkable variety of cell types which interact physically and chemically in order to provide function. Diseases are ecosystems out of balance, but to rebalance them we need to understand how they are built and how they work. To unlock this problem, we:
Map human tissues from development across the human lifespan and in disease cohorts using single cell and spatial multiomics. We will build comprehensive tissue atlases that serve as foundational resources for the field in the same way that genome sequences do. We will understand how the human body is built and characterise diseases at unprecedented resolution.
Perturb human tissues in vitro using advanced multilineage organoid systems, allowing us to test the predictions of the atlases, to model tissues and specific diseases, and to test how population-scale genetic variation impacts tissue ecosystem function.
Model human tissues in silico using artificial intelligence to reveal hidden biological associations from the in vivo atlases and in vitro perturbation studies and perform theoretical experiments, accelerating our capacity to test biological mechanisms at scale.
These three approaches are integrated into a “lab-in-the-loop” workflow – with foundation models continually guiding our mapping and perturbations.
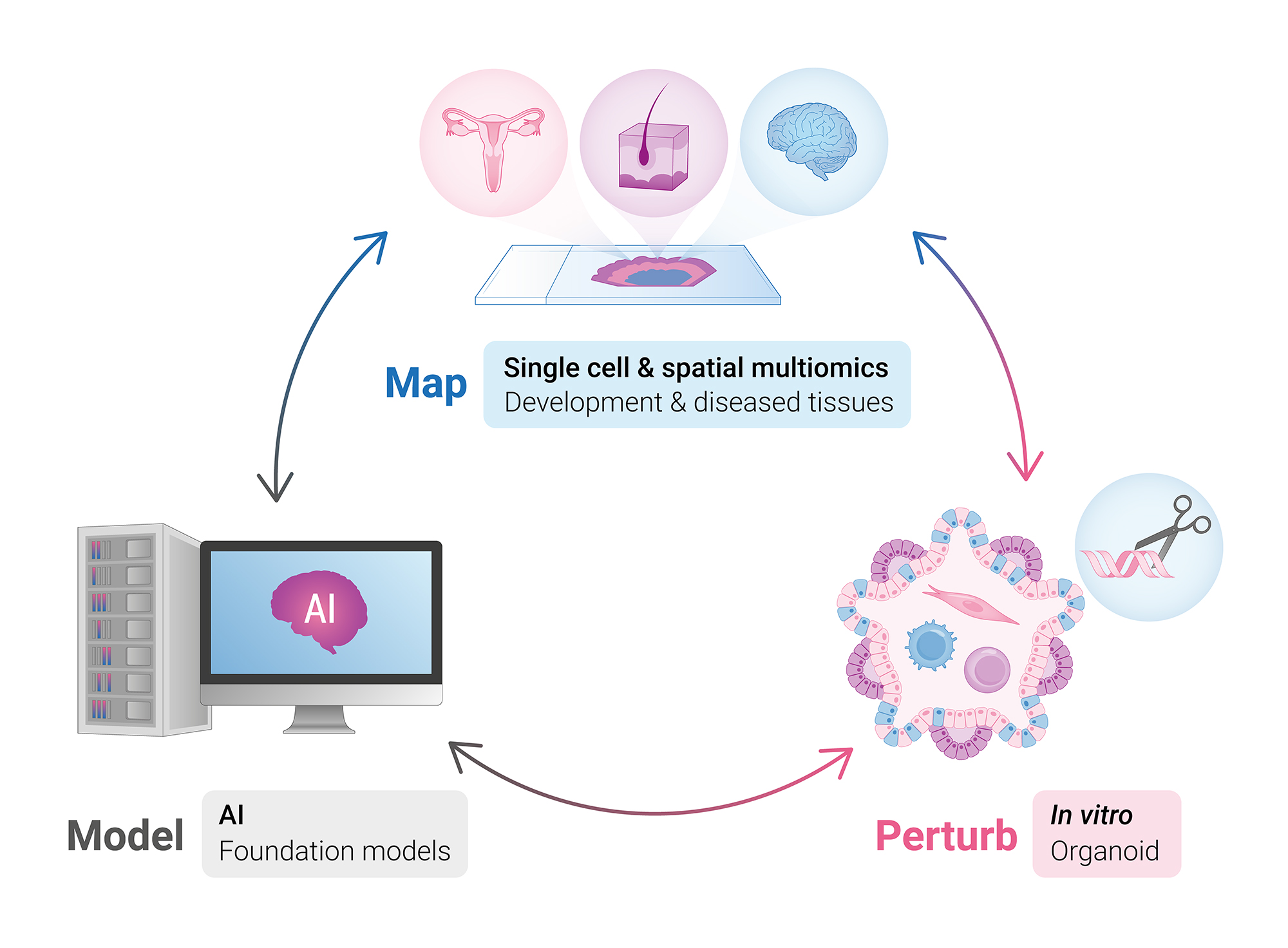
The Cellular Genomics Programme’s unique merging of massive multimodal developmental and clinical data with AI/ML will help us close the gap between genotype and phenotype in biomedicine – enabling an end-to-end understanding of disease for patient benefit and new strategies for tissue engineering.
Integrated multidisciplinary expertise
Our group leaders are world-leading scientists and clinician-scientists with wide ranging expertise including in the cellular basis of immune disorders, the molecular drivers of childhood cancers, and the development and function of reproductive tissues and the brain. We are a centre for technology development, both in terms of hardware (e.g. spatial genomics, multiomics) and software (e.g. foundation models, bioinformatic tools). Our scientists are key contributors to the Human Cell Atlas international consortium, which aims to create comprehensive reference maps of all human cells.
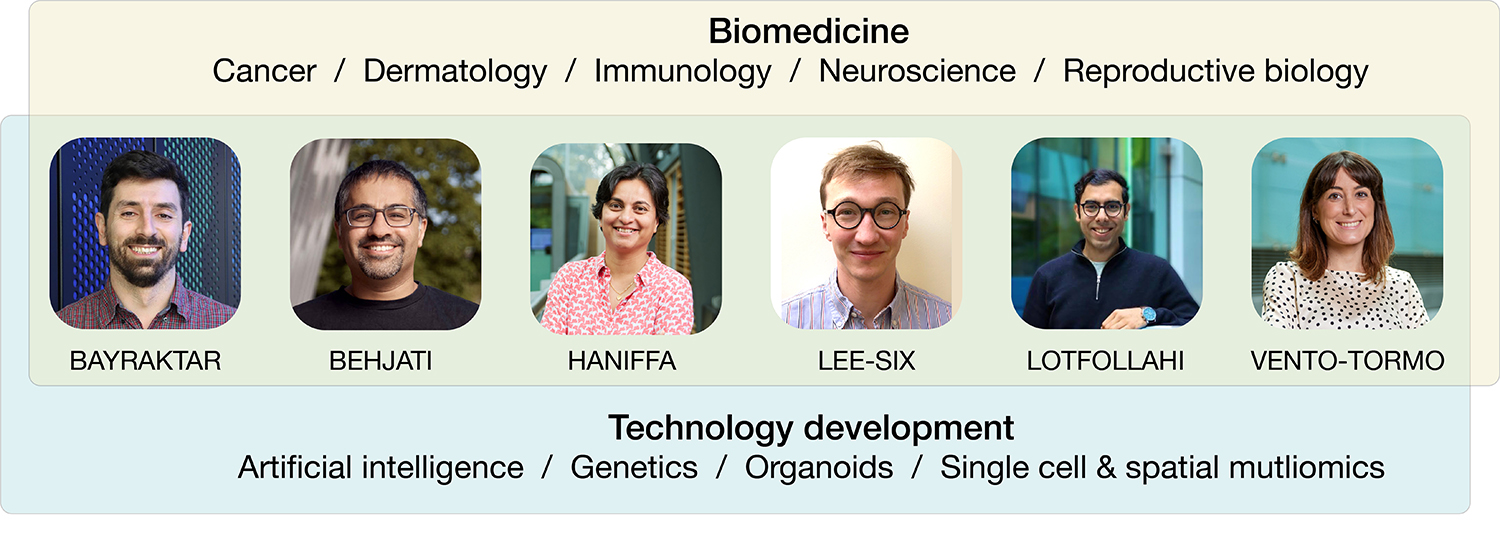

In addition to our six Faculty, our four Associate Faculty colleagues bring leading expertise in immunology, spatial techniques and AI/ML to help us meet our vision.

Our success
Publications
Cellular Genomics researchers publish widely impactful papers in human tissue atlasing and disease genomics. Our recent highlights include:
Spatiotemporal cellular map of the developing human reproductive tract. Led by Valentina Lorenzi and Cecilia Icoresi-Mazzeo. Vento-Tormo Lab. Nature.
A non-canonical lymphoblast in refractory childhood T-cell leukaemia. Led by Bram Lim and Holly Whitfield. Behjati Lab. Nature Communications.
A single-cell and spatial genomics atlas of human skin fibroblasts reveals shared disease-related fibroblast subtypes across tissues. Led by Lloyd Steele. Haniffa Lab. Nature Genetics.
Quantitative characterization of cell niches in spatially resolved omics data. Led by Sebastian Birk. Lotfollahi Lab. Nature Genetics.
Integrated histopathology, spatial and single cell transcriptomics resolve cellular drivers of early and late alveolar damage in COVID-19. Led by Jimmy Lee and Sam Barnett. Bayraktar Lab. Nature Communications.
We are also committed to open science and post our papers in advance on bioRxiv. Our recent uploads include the Lotfollahi lab’s tools for predicting cellular perturbations and modelling gene programmes, the Haniffa lab’s exploration of tissue niches in the skin and cell atlas of prenatal human development, the Bayraktar lab’s spatial atlases of autism-associated genes and glioblastoma, and the Vento-Tormo lab’s multiomic atlas of B cell development and tool to predict metabolic activity in single cell data.
Prizes
The impact and success of our science has been recognised by the scientific community, as shown in this selection of recent awards.