Respiratory Virus and Microbiome Initiative
Parasites and Microbes
Aims
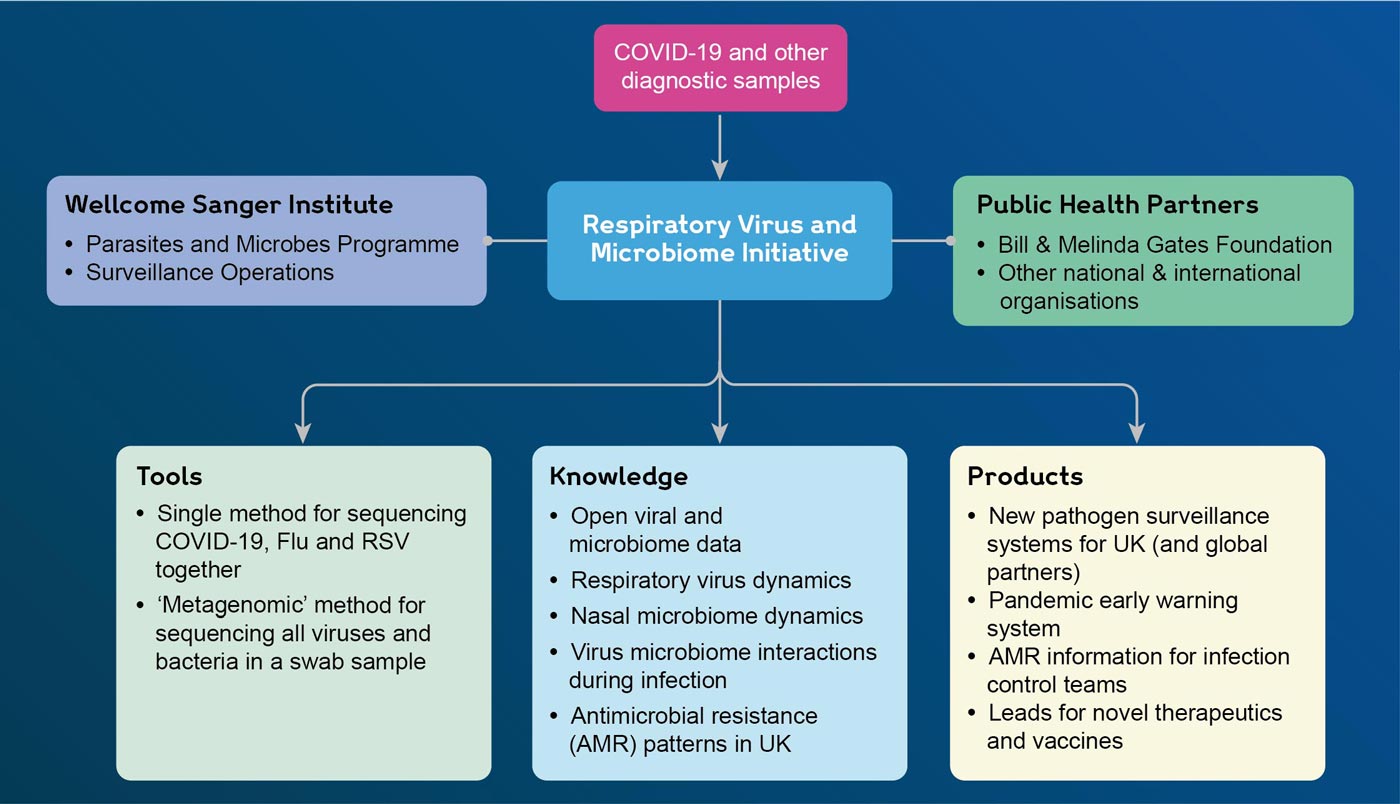
The Respiratory Virus and Microbiome Initiative aims to establish capabilities for large-scale genomic surveillance of respiratory viruses such as influenza virus and respiratory syncytial virus (RSV), and to survey for as yet unknown pathogens. By establishing standardised approaches for combined sequencing of SARS-CoV-2, and other common respiratory viruses in a single assay and developing pathogen-agnostic sequencing approaches, the Initiative will enable:
- Generation of knowledge on the transmission and evolution of known respiratory viruses and other pathogens.
- Discovery of novel respiratory viruses to inform early warning systems for new pandemic threats.
- Deeper understanding of respiratory microbiome dynamics and how this influences health and infection.
- Tracking and understanding of antimicrobial resistance in the respiratory tract at a population level.
Revealing co-infections with metagenomics
Co-infection and secondary infections caused by bacterial and fungal pathogens are common in severe respiratory viral infections, including for SARS-CoV-2, RSV, and pandemic influenza. As such, a major area of focus for the Respiratory Virus and Microbiome Initiative is on developing the tools and approaches needed to use shotgun metagenomics for untargeted surveys of the microbial diversity present in respiratory swab samples. These approaches will enable a better understanding of how the respiratory microbiota changes during infection, how this relates to variability in the severity of illness between individuals, and can identify opportunities to intervene earlier to prevent or blunt a respiratory pathogen.
Metagenomics also has the potential to allow tracking of antimicrobial resistance (AMR) in the respiratory microbiome, which will be increasingly important in the years to come as the arsenal of effective antimicrobial dwindles.
About the Respiratory Virus and Microbiome Initiative
 The Initiative is uniquely placed to undertake this work:
The Initiative is uniquely placed to undertake this work:
- Our team has gained deep experience from undertaking the nationwide genomic surveillance for COVID-19 as part of the COVID-19 Genomics UK (COG-UK) consortium and now in collaboration with the UK Health Security Agency (UKHSA).
- The Sanger Institute has a long history of developing cutting-edge genomic techniques and innovative sequencing approaches.
Research and development work undertaken by the Respiratory Virus and Microbiome Initiative will both benefit from, and contribute to, the core aims of the Sanger Institute’s Parasites and Microbes programme, and public health.
All data, protocols and methods generated through the Initiative will be made freely and publicly available. Over time the platforms and technologies developed by Initiative will be scaled up and productised through the Sanger Institute’s Genomic Surveillance Unit so that public health partners can adopt them as part of their pathogen genomic surveillance programmes.
Join our team
There are a range of exciting projects underway in our team. We are always on the lookout for bright, motivated people to join the group so, if you’re interested, please drop Ewan Harrison an email and include your CV.
Acknowledgements
The work of the Respiratory Virus and Microbiome Initiative is supported through interactions with the following people:
- Dinesh Aggarwal
- Tobi Ajenifuja
- Camilo Gomez Arenas
- Cristina Ariani
- Katherine Auger
- Alp Aydin
- Karen Baker
- Kristina Battleday
- Mathew Beale
- Tim Beard
- Graham Beaver
- Katie Bellis
- Emma Betteridge
- Beth Blane
- Roisin Boggan
- Joe Brennan
- Tanya Brooklyn
- Hilary Browne
- Josie Bryant
- Victoria Carr
- Peter Clapham
- Jaime Tovar Corona
- Jo Crofton-Diggins
- Bruhad Dave
- Matthew Dorman
- Jillian Durham
- Sarah Francis Field
- Katherine Figueroa
- Emma Fraser
- Anastasia Galvin
- Sonia Goncalves
- Marina Gourtovaia
- Julia Harvison
- Kevin Howe
- Ya-Lin Huang
- Chrisa Humphreys
- Stephen Inglis
- Ali Jabir
- David Jackson
- Wiesia Johnson
- Ian Johnston
- Amie Jordi
- Keely Kay
- Sally Kay
- Jon Keatley
- Alan Keith
- Marissa Knoll
- Prasanth Kothuri
- Cordelia Langford
- Florent Lassalle
- Adrianne Lignes
- Rich Livett
- Rhona Long
- Lerato Magosi
- Antonio Marinho
- Ben McCarthy
- Duncan Ng
- Fernanda Novaes
- Katherine Oswald
- Naomi Park
- Maggie Payne
- Martin Pollard
- Tom Preston
- Stacey Price-Weight
- Michael Quail
- Diana Rajan
- Sarah Reeks
- Joe Reynolds
- Joe Robinson
- Eduardo Martin Rojo
- Jeff Ryen
- Beth Sampher
- Frank Schwach
- Carol Scott
- Olivier Seret
- Yan Shao
- Lesley Shirley
- Maqsood Siddiqui
- Carol Smee
- Catarina Sousa
- Andrew Sparkes
- Sara Stott
- Alyce Taylor-Brown
- Diego Teixeira
- Nashma Thesin
- Nick Thomson
- Wendy Thorpe
- Thomas Thron
- Scott Thurston
- Josef Wagner
- Jerry Watts
- Thomas Whiteley
- Andrew Wong
- Michael Woodward
- Ruth Wright
- Rory Yeadon
RVI Scientific Advisory Board
- Professor Emma Thomson, MRC-University of Glasgow Centre for Virus Research
- Professor Judith Breuer, University College London (UCL), Division of Infection and Immunity
- Dr Meera Chand, Public Health England and Consultant with Guy’s and St Thomas’ NHS Foundation Trust
- Dr Josefina Campos, Director of the National Genomics and Bioinformatics Center at ANLIS Malbrán
- Professor Thushan De Silva, University of Sheffield, Department of Infection, Immunity and Cardiovascular Disease
Scientific Collaborations
The Respiratory Virus and Microbiome team is working in close collaboration with David Bonsall [well.ox.ac.uk] from the Wellcome Centre for Human Genetics and Pandemic Sciences Institute, University of Oxford, and Christophe Fraser [bdi.ox.ac.uk] Pandemic Sciences Institute and Big Data Institute, University of Oxford to develop low cost metagenomic sequencing methods and analysis tools suitable for global use.
Core team

Katie Bellis
Staff Scientist

Dr Róisín Boggan
Postdoctoral Fellow

Dr Wendy Figueroa
Postdoctoral Fellow

Dr William L. Hamilton
Clinical Lecturer in Medical Microbiology

Dr Marissa Knoll
Postdoctoral Fellow

Dr Duncan Ng
Postdoctoral Fellow

Dr Ouli Xie
Postdoctoral Fellow (ESPOD)
Related groups
Partners
We work with the following groups
