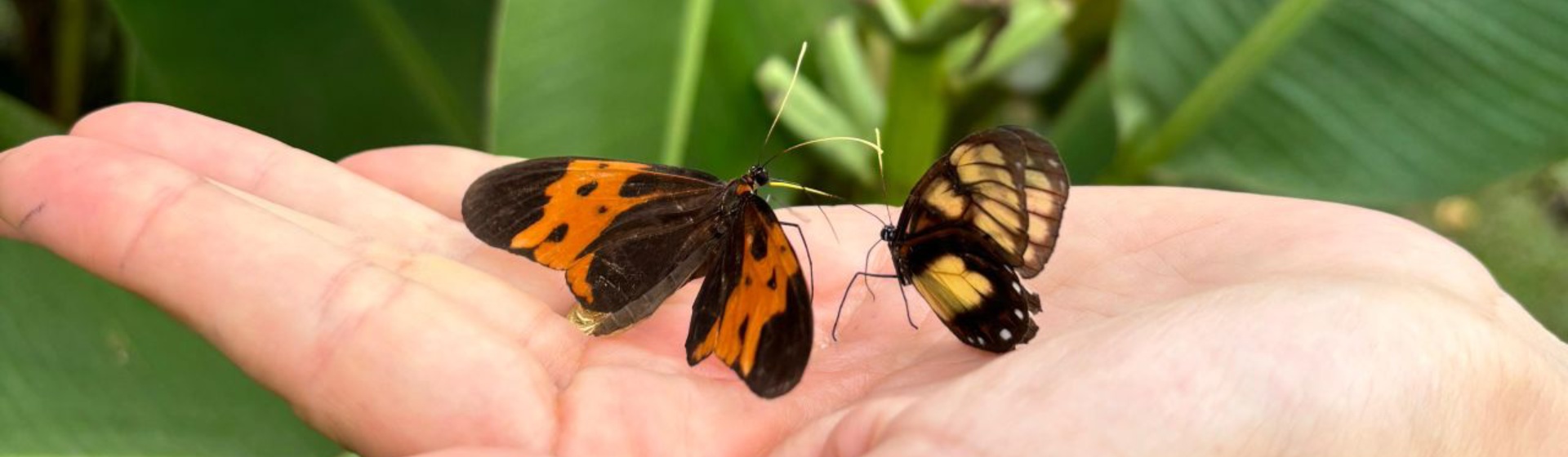
Tree of Life
Overview
Species collected
12,500
Species collected for DNA sequencing
Species submitted
3,949
Full species’ genome sequences shared with the global genomics research community
Sequencing total
2,458,722
Tbp read so far
Celebrating 3,000 genomes
The Tree of Life Programme uses DNA sequence to investigate the diversity and origins of life on Earth. We focus on generating rich data for all eukaryotic organisms, which means all complex life with a nucleus in their cells – that is every animal, plant, fungi and protist on the planet.
At the core of our research is the production of high-quality reference genomes for individual species, which our teams are producing at an unprecedented scale. Alongside this reference genome production, research teams within Tree of Life are exploring questions around ecosystem function, species radiations, reproductive diversity and more.
Tree of Life’s projects operate in tandem with the Earth BioGenome Project. This global network of over 60 affiliated biodiversity genomics projects shares the common aim to “sequence life for the future of life”. Our overarching goal is to produce reference genomes for all 1.6 million known species on Earth, and make this library of knowledge accessible to the world for conservation, biomonitoring, bioindustry and better understanding life on our shared planet. All our data – including as much work-in-progress as possible – is published openly and available freely to researchers across the world.
What we do
We produce chromosomally-complete reference genomes for all our target species. A chromosomally-complete genome is one where very nearly all the genome has been ordered and oriented into what we believe are representations of the chromosomes of the species. Ensuring the genomes are as complete as we can make them means that users of the genomes can be assured they have the highest-quality data as the foundation of their work.
Why reference genomes?
Tree of Life has assembled a world-leading genome production pipeline – the Genome Engine. The Genome Engine process assures the highest quality across our work, from the ethics and legal status of collections, through shipping, lab processing, sequencing, assembly, curation and release.
To find out more about our processes, please see our Genome Engine page.
How do we produce our genomes?
As well as leading or participating in major collaborative projects, Tree of Life Faculty, Associate Faculty, Honorary Faculty and our postdoctoral fellows and scientific staff use biodiversity genomics approaches across a wide range of topics, from ongoing speciation to deep phylogenetics, from within-species diversity to wide-ranging surveys of genomic novelties, and including development of tools and standards for biodiversity genomics.
| Speciation | How and why do species radiations occur? | Mara Lawniczak | Joana Meier | Richard Durbin |
| Chromosome evolution | What drives chromosomal change (or stasis) in diverse major lineages? | Joana Meier | Mark Blaxter | Kamil Jaron |
| Non-Mendelian inheritance | How have various forms of “cheating” of the Mendelian rules evolved and how do they impact species’ biology? | Mark Blaxter | Kamil Jaron |
| Genomics of parasitism | What do parasites do differently? | Mark Blaxter | Mara Lawniczak |
| The microbiomes of eukaryotes | What do the microbial genomes sequenced accidentally along with their hosts tell us about patterns of symbiosis? | Mark Blaxter |
| The evolution of sexual systems | Why are some species simply male-female, while others have a wide range of reproductive modalities? | Kamil Jaron |
| The origin and evolution of sex chromosomes | What distinguishes sex chromosomes? How do they arise and how do they degrade and disappear? | Kamil Jaron | Joana Meier | Mark Blaxter |
| Building phylogenies from genomes | Is there a “tree of life” does rampant hybridisation and introgression mean we should rather think of it as a “net of life”? | Richard Durbin | Joana Meier | Mark Blaxter |
| Better software, better genomes | Can we assemble genomes better by developing new AI-informed tools and processes? | Kamil Jaron | Richard Durbin | Gene Myers | Mark Blaxter |
| The uniqueness of Bats | What genomic features underpin the extraordinary biology (flight, echo location, longevity, virus resistance) of bats? | Emma Teeling |
| The origin of cell types across diversity | Are cell types (“skin”, “gut”, etc) homologous across animals, and when did they arise and how are they determined? | Arnau Sebé-Pedrós | Mara Lawniczak | Mark Blaxter |
Tree of Life research
The Tree of Life Programme is a partner in many research projects (see also below under ‘Collaborations’).
 Darwin Tree of Life (2019- ongoing)
Darwin Tree of Life (2019- ongoing)
The aim of the Darwin Tree of Life (DToL) project is to produce reference genomes for each of the estimated 70,000 eukaryotic species in Britain and Ireland.
DToL is a partnership between biodiversity, genomics and analytics partners: Sanger, the Earlham Institute, EMBL-EBI, the Marine Biological Association, the Natural History Museum in London, the Royal Botanic Gardens Edinburgh, the Royal Botanic Gardens at Kew, CABI, the Institute of Zoology, and the universities of Cambridge, Edinburgh, Oxford and Dublin.
For progress in data generation in DToL, see https://goat.genomehubs.org/projects/DTOL.
To find out more, visit the Darwin Tree of Life website. Or follow DToL on Twitter @darwintreelife.
 Aquatic Symbiosis Genomics (2020-2025)
Aquatic Symbiosis Genomics (2020-2025)
The aim of the Aquatic Symbiosis Genomics (ASG) project is to better understand symbiosis through sequencing of the “host” and the microbial symbionts in a wide range of species that live in the sea or freshwater. ASG is jointly funded by the Wellcome Sanger Institute and the Gordon and Betty Moore Foundation, with ten global partners acting as hubs for different groups of symbiotic organisms.
For progress in data generation in ASG, see https://goat.genomehubs.org/projects/ASG.
To find out more, visit the Aquatic Symbiosis Genomics website.
 BIOSCAN in the UK (2020-ongoing)
BIOSCAN in the UK (2020-ongoing)
Tree of Life’s BIOSCAN UK project aims to study the genetic diversity of 1,000,000 flying insects from across the UK. Insects from 100 sites are being collected on a monthly basis over five years by project partners and then analysed at Sanger using DNA barcoding. The resulting sequence data will provide a baseline characterisation of insect species diversity over space and time and thus form a much needed resource for DNA-based biomonitoring in the UK. BIOSCAN UK is affiliated to the IBOL-Europe stream of the Biodiversity Genomics Europe (https://biodiversitygenomics.eu/) Horizon programme of the EU.
To find out more, visit the BIOSCAN webpage.
 Project Psyche (2024-ongoing)
Project Psyche (2024-ongoing)
Project Psyche is a multinational effort to sequence the genomes of all 11,000 species of Lepidoptera – moths and butterflies – in Europe. Currently it is in phase 1, where we aim to sequence 2000 species and all specimens are provided by seven sampling hubs across Europe and sequencing and assembly of the genomes is performed at the Sanger Institute. We are currently working on publications using the first 1000 genomes.
For progress in data generation in Psyche, see https://goat.genomehubs.org/projects/Psyche.
To find out more, including how to join the Psyche community, visit the Project Psyche website.
 AEGIS (2024-ongoing)
AEGIS (2024-ongoing)
The Ancient Environmental Genomics for Improving Sustainability programme is a global initiative centred on Copenhagen and Cambridge that brings together geologists, archaeologists, ancient DNA specialists and genomics to better understand the response of ancient ecosystems to climate change. Funded by the NovoNordisk Foundation and the Wellcome Trust, the seven-year project aims to identify patterns of change in the genetics of ancient ecosystems, derived from sequencing of aeDNA from sediment cores going back hundreds of thousands of years, and to engineer those changes into modern crops to improve sustainable production. Tree of Life is delivering genome reference sequences of relevant modern species. See the AEGIS website (https://globe.ku.dk/research/ancient-environmental-genomics-initiative-for-sustainability/) for details.
 The European Reference Genome Atlas (ERGA) (2022-2025, and ongoing)
The European Reference Genome Atlas (ERGA) (2022-2025, and ongoing)
The European Reference Genome Atlas is a continental initiative to build genomic resources and activity across Europe, and to generate reference genome sequences for European biodiversity. Tree of Life was part of the ERGA Pilot Project, and a participant in the Biodiversity Genomics Europe (https://biodiversitygenomics.eu/) Genome Stream project funded by the Horizon initiative and the UK government.
For progress in data generation in ERGA, see: https://goat.genomehubs.org/projects/ERGA.
See the ERGA website for further details: https://www.erga-biodiversity.eu/.
 The Tree of Sex (2023-ongoing)
The Tree of Sex (2023-ongoing)
The Tree of Sex is a global collaboration that aims to define, describe and analyse the many different modes of reproduction present in the world of Eukaryota, and to use these data to explore the impacts and evolution of reproductive mode across the tree of life. Tree of Life is providing core informatics support for development and deployment of the Tree of Sex database (see https://www.treeofsex.org/.
 The Vertebrate Genomes Project
The Vertebrate Genomes Project
The VGP was founded to sequence the genomes of all ~70,000 species of fish, reptiles, birds, mammals and allies. Researchers at the Sanger have been part of the Vertebrate Genomes Project since its inception over a decade ago. We contribute new genomes to the growing roster of high-quality assemblies available, and are working with VGP colleagues round the world in analysing the “Phase 1” data freeze, which includes 500 genomes from across vertebrate diversity.
For progress in data generation in the VGP, see https://goat.genomehubs.org/projects/VGP.
See https://vertebrategenomesproject.org/.
 The Earth BioGenome Project
The Earth BioGenome Project
Tree of Life faculty are also engaged with many additional international collaborative projects, especially those under the banner of the Earth BioGenome Project.
See the EBP website (https://www.earthbiogenome.org/) for details of these initiatives and the EBP overview on Genomes on a Tree at https://goat.genomehubs.org/projects/EBP.


