Open Targets
Overview
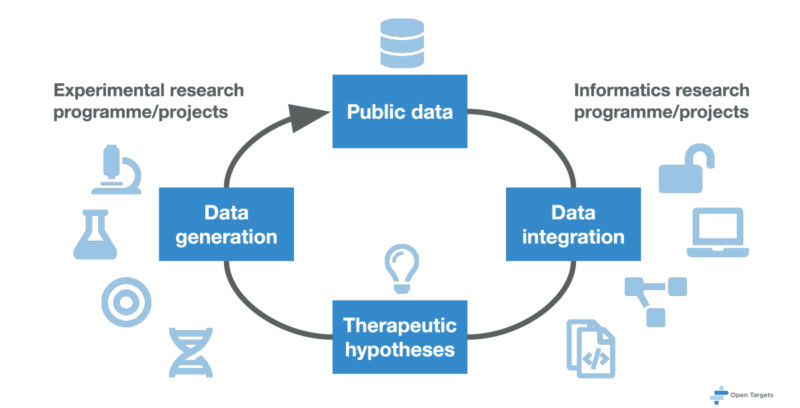
Open Targets offers an early assessment of the expected efficacy of a therapeutic intervention for a given drug target, with the aim of improving the almost 90 per cent failure rate of drug discovery efforts in clinical trials. The extensive programme of experimental and informatics projects uses large-scale experiments, analysis, and cutting edge informatics technologies to provide a more complete understanding of the biological drug target. Our projects are designed to identify causal links between targets and disease in three focus areas: oncology, neurodegeneration, and immunology and inflammation, as well as provide disease agnostic informatics frameworks for target identification and prioritisation.
Our research projects have made significant contributions to understanding the relationships between targets and diseases, presented in our series of high-impact research publications.
Details on our resources can be found on our website.