Production Software Development
Laboratory Information and Management Systems (LIMS) compute and infrastructure
How we support Sanger’s science
Our work enables our researchers to carry out large-scale, high-throughput experiments across a vast range of research areas and experimental approaches. To achieve this, we use a combination of bespoke software we have written ourselves and commercial software solutions that we support.
While our scientists’ work covers probing cancerous cells, monitoring the spread of bacteria and viruses through populations, and genomically identifying the full biodiversity of the insects captured in a sample, many of their needs are the same. All the samples that come to the Sanger Institute need to be logged, stored, retrieved, experimented on, and sequenced. And this data needs to be easily created, used, and accessed.
Our solutions record everything about where the sample comes from, what it is, and what the legal requirements around it are. This is then combined with details about any experiments that are carried out on the sample before it is sent through for DNA sequencing. We then combine the experimental and sample data with the genomic data coming out of the sequencing machines so that our scientists can carry out their analysis to gain new insights into health and disease, human development, and biodiversity.
Our Approach
Our approach is to maximise the value we deliver while minimising the work required. We work closely with our end users to understand their requirements and identify the most effective solutions, which can often involve no additional software whatsoever.
We allocate time to continuously address technical debt and aim to develop and maintain quality solutions. To power further research across the wider genomic scientific community, our software is open source and freely available on github.
Streamlining and standardising laboratory management processes
To support the Institute’s science, we work closely with research groups across all the research programmes, and the Scientific Operations teams to modernise and digitize our laboratory processes, workflows, and data harvesting. This work will enable greater streamlining and reproducibility of the Institute’s science.
COVID-19 monitoring and surveillance
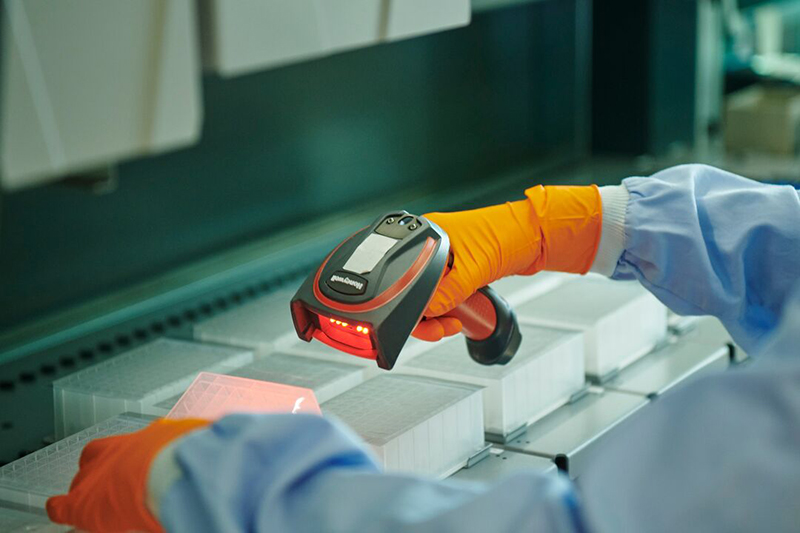

We played an integral role in the set up and running of the systems and software used by the Sanger Institute to deliver nationwide genomic surveillance of COVID-19 in the UK for the Government to monitor and track the rise and spread of new variants of the virus.
This work involved building systems to log, store, track, move and prepare 65,000 samples a week as they were processed by the Institute.
Biodiversity monitoring
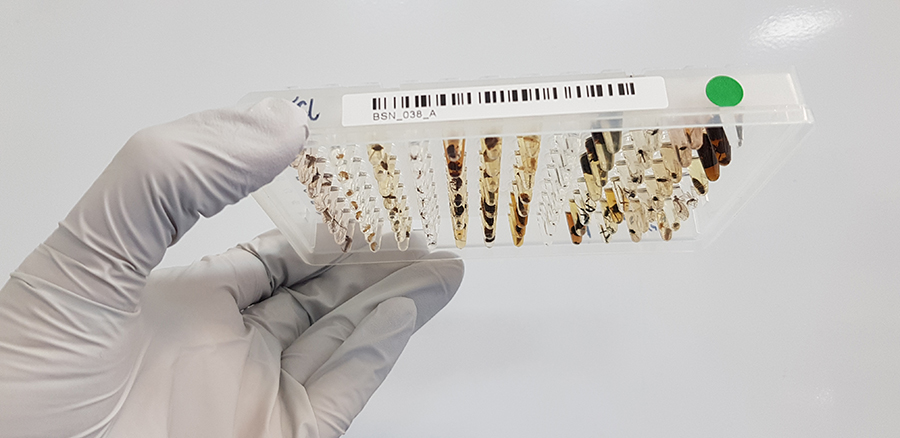
We are closely involved in integrating the robotic automation required for the BIOSCAN project into our systems for our Tree of Life scientists. In this work, up to 10,000 different insects are being put into a single well for DNA sequencing and subsequent identification.
Not many things like having 10,000 insects inserted into them, and scientific data systems are no exception. To enable this work, we are helping to develop and run the robotics and tagging processes required.
Respiratory health

We are also working closely with the Sanger Institute’s researchers in the Respiratory Virus and Microbiome Initiative to help them build the workflows and supporting systems needed to log, store, prepare and sequencing tens of thousands of samples.
Spatial-Omics
Another key area that we are helping to develop is the Sanger Institute’s spatial-omics work where fine-grained imaging at vast scale is being combined with DNA sequencing to understand the workings of single cells within the tissues of the body. Our work aims to connect all the data from tracking the samples at delivery and experimentation, through DNA sequencing and imaging so that it can be combined to give the full view of how cells function within the body.
How we work
Our team draws from a wide range of experience and knowledge including Computer Science and Biology. We work collaboratively with our research partners and within the team to develop the best possible solutions. We value diversity of ideas and knowledge and seek to grow our team members to their full potential. We continually develop our professional learning through courses, conferences, and dedicated learning sessions. A number of the team have obtained Masters and part-time Open University degrees as part of their career development.
Recruitment
We are always looking for talented individuals to join the team. These roles can be found in the Current Vacancies area of the Careers section of the website, and are advertised on Twitter – @Sangercareers
Core team

Ene Göktan
Informatics & Digital Associate

Stephen Inglis
Senior Software Developer - Production Software

Seena Nair
Senior Software Developer

Neil Sycamore
Bioinformatician

Katy Taylor
Senior Software Developer

Wendy Yang
Senior Software Developer
Previous core team members

Beth Flint
Senior Scientific Manager

Dr James Glover
Senior Software Developer

Rich Livett
Technical Delivery Lead (Platform and software development)
