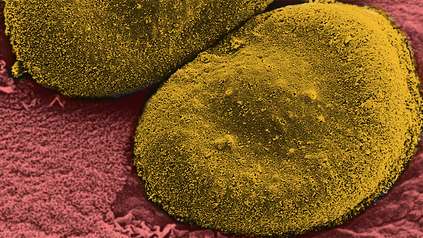
Pneumonia mapped in largest genomic survey of any disease-causing bacterium
The research is published in The Lancet Infectious Diseases today (10 June), and with a sister paper in EBioMedicine, reveals which strains of S. pneumoniae (also known as the pneumococcus) are circulating around the world and explains why pneumococcal pneumonia rates are still high despite the existing vaccines. Funded by a grant from the Bill & Melinda Gates Foundation, this work will help predict which strains will be important for new pneumococcal vaccines, and shows that ongoing global genomic surveillance is vital.
Many countries around the world have introduced the pneumococcal conjugate vaccine (PCV)** over the last ten years. This vaccine, which targets the coat around each S. pneumoniae bacterium, has greatly reduced the number of childhood infections. However, while PCV is highly effective against up to 13 important coat types, there are over a hundred types known, and despite the vaccine, pneumococcal pneumonia rates remain very high.
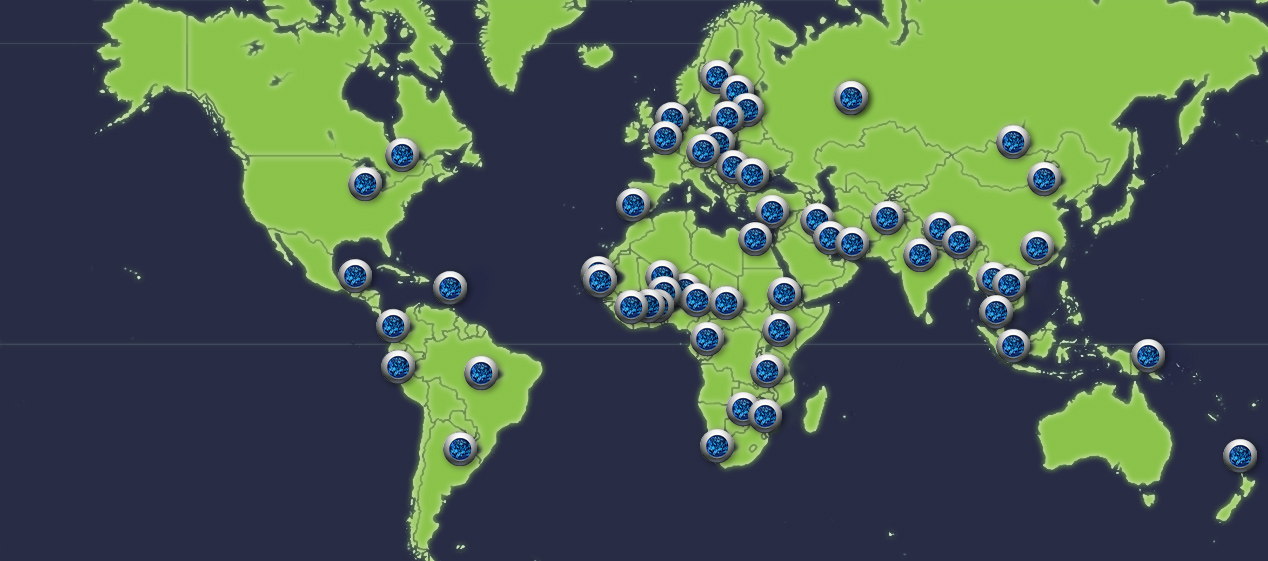
To understand and help combat this infection, researchers set up the Global Pneumococcal Sequencing project (GPS) to carry out genomic surveillance of S. pneumoniae worldwide. Working with partners around the world, the researchers sequenced the DNA of over 20,000 S. pneumoniae samples from infected people from 51 countries.
Samples were collected both before and after PCV introduction, and the DNA sequences and health data were compared. This makes it possible to determine changes in the bacteria that could affect how well the vaccine protects against the pneumococcus, and whether new strains are emerging that would impact disease severity and ease of treatment.
The researchers discovered 621 genetic strains globally, each associated with one or more coat types. They also saw that the levels of non-vaccine type bacteria rose after the introduction of PCV, showing how bacteria evolve in response to the vaccine.
“Pneumonia is a huge threat to health worldwide. We now have an unprecedented view of the global population of S. pneumoniae bacteria, the usual cause of bacterial pneumonia, and can see evolutionary changes that lead to vaccine evasion. This will give crucial information for future vaccine strategy worldwide, and help save lives.”
Professor Stephen Bentley Senior author on the papers, from the Wellcome Sanger Institute
“Our study gives the first genomic description of the S. pneumoniae population of the world. This has never been possible before, as previously only samples from individual populations had been studied. Now we have global data, showing which strains are present in each country, and can use this to understand pneumococcal infection on a world-wide scale.”
Dr Rebecca Gladstone Joint first author on two of the papers, from the Wellcome Sanger Institute
The pneumococcus can cause disease in other areas of the body too, for example infecting the brain or blood, causing meningitis or bloodstream infections, which can all lead to sepsis. Infant vaccination with PCV protects against these pneumococcal infections too. By reducing the transmission of S. pneumoniae between children, PCV also reduces the number of adult infections through herd immunity.
“It is vital to understand the strains of S. pneumoniae present around the world, and how they respond to the introduction of PCV. This genomic study allows us to not only see the current most important global strains but also helps understand their evolution. This information will open the door to developing predictive tools to identify the strains likely to emerge in response to vaccine use.”
Dr Lesley McGee Co-Principal Investigator on the project from the Streptococcus Laboratory at the Centers for Disease Control and Prevention in the United States
“GPS turns a spotlight onto a new era in which the intersection of genomics and public health enables unparalleled capacity for optimizing prevention strategies, while providing an immensely valuable tool for forecasting and addressing new challenges ahead.”
Professor Robert Breiman Director of the Emory Global Health Institute, and Principal Investigator for the project
“We must continue to immunize children around the world because vaccination is the single best way of reducing the risk of pneumonia, as it prevents children passing S. pneumoniae between them and adults. However, we are fighting a battle against evolution of bacterial strains. This research shows the importance of ongoing global genomic surveillance to understand which strains are likely to cause a threat, to help reformulate the next generation of vaccines.”
Professor Keith Klugman Director of the Pneumonia team at the Bill & Melinda Gates Foundation
More information
* Pneumonia statistics from the World Health Organization:
Pneumonia is the single largest infectious cause of death in children worldwide. Pneumonia killed 920 136 children under the age of 5 in 2015, accounting for 16% of all deaths of children under five years old.
https://www.who.int/news-room/fact-sheets/detail/pneumonia
**PCV vaccine.
The pneumococcal conjugate vaccine, PCV, targets up to 13 specific types of the polysaccharide coat of the S. pneumoniae, and works extremely well for those specific types. However, there are about 100 different types of polysaccharide coat so it is vital to predict the most important type. It is generally used to vaccinate children under the age of two years.
https://www.nhs.uk/conditions/vaccinations/pneumococcal-vaccination/
Publications:
- Stephanie Lo & Rebecca Gladstone et al. (2019) Pneumococcal Lineages Associated with Serotype Replacement and Antibiotic Resistance in Childhood Invasive Pneumococcal Disease in the Post-PCV13 Era: An international whole genome sequencing study. The Lancet Infectious Diseases. DOI: 10.1016/S1473-3099(19)30297-X
- Rebecca Gladstone & Stephanie Lo et al. (2019) International Genomic Definition of Pneumococcal Lineages to Contextualise Disease, Antibiotic Resistance and Vaccine Impact: A Whole Genome Bacterial Sequencing Study. EBioMedicine. DOI: 10.1016/j.ebiom.2019.04.021
Funding:
These studies were funded by the Bill & Melinda Gates Foundation, the Wellcome Sanger Institute and the U.S. Centers for Disease Control and Prevention.
Selected websites
The Global Pneumococcal Sequencing Project
The Global Pneumococcal Sequencing Project is a worldwide genomic survey of the impact of vaccination on the population of Streptococcus pneumoniae. https://www.pneumogen.net/gps/
Bill & Melinda Gates Foundation
Centers for Disease Control and Prevention
CDC works 24/7 to protect the health, safety, and security of Americans. CDC fights disease whether it starts at home or abroad, is infectious or not, occurs naturally, by accident, or from a deliberate attack. CDC promotes the health and well-being of Americans of all ages—doing all it can to prevent infections, injuries, and illnesses from ever occurring. https://www.cdc.gov/
Emory University
Emory University is known for its demanding academics, outstanding undergraduate experience, highly ranked professional schools and state-of-the-art research facilities. Emory encompasses nine academic divisions as well as The Carter Center, the Emory Global Health Institute, the Yerkes National Primate Research Center, the Michael C. Carlos Museum, and Emory Healthcare, Georgia’s largest and most comprehensive health care system. http://www.emory.edu
The Wellcome Sanger Institute
The Sanger is one of the world’s leading genome and biodata institutes. Through its ability to conduct research at scale, it is able to engage in bold and long-term exploratory projects that are designed to influence and empower science globally. Institute research findings, generated through its own research programmes and through its leading role in international consortia, are being used to develop new diagnostics and treatments for human disease and to understand life on Earth. Find out more at www.sanger.ac.uk or follow @sangerinstitute on Twitter, Facebook, LinkedIn and on our Blog
About Wellcome
Wellcome exists to improve health by helping great ideas to thrive. We support researchers, we take on big health challenges, we campaign for better science, and we help everyone get involved with science and health research. We are a politically and financially independent foundation.