About
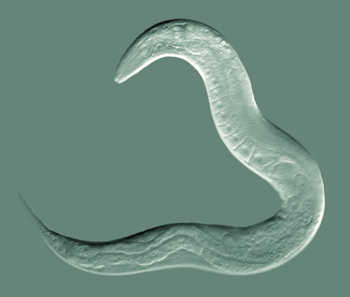
Caenorhabditis elegans (informally known as ‘the worm’) is a small soil-dwelling nematode that is widely used as a model system for studies of metazoan biology. C. elegans‘ popularity results from the confluence of several factors: its developmental program is understood at the single-cell level; it is highly amenable to genetic manipulation, including RNAi intervention; and it has a complete, high-quality reference genome sequence.
Genomic data for C. elegans, C. briggsae and a host of other nematodes can be found at WormBase.
Background
Caenorhabditis genome sequencing
C. elegans was the first animal to have its genome completely sequenced. The WTSI’s contribution to this effort was significant. Indeed The project was one of the flagship activities in the early life of the WTSI, and as such is one of the defining legacies of the institute itself.
The mapping and sequencing of the reference genome was a joint project between The Wellcome Trust Sanger Institute and The Genome Institute at Washington University (St. Louis). The essentially-complete sequence was formally published in December 1998, and data was made regularly and freely available in advance of publication. The last remaining gap was closed in 2002, although the genome sequence continues to be scrutinzed and improved as new evidence is published.
In addition to C. elegans, The WTSI and the WUGI also collaborated on the genome sequencing of the related nematode
Caenorhabditis briggsae. A whole-genome-shotgun assembly was made available in July 2002 and formally published in November 2003.
WormBase
WormBase is a collaborative project to capture Curate and distribute information about C. elegans biology. It began life as ACeDB, a database application software package developed by jointly Richard Durbin at the Sanger Institute and Jean-Thierry Mieg. ACeDB was used extensively during the course of the
C. elegans sequencing project to coordinate the sequencing effort and to integrate the worm sequence with the genetic and physical maps.
WormBase was originally started in 2000 as a way to make data in ACeDB easily accessible via a web-browser. From the outset The project was heavily committed to the curation and interpretation of the C. elegans literature, and rapidly moved from a genome-centric perspective to one that more evenly balances the worm genome with other aspects of its biology.
The original WormBase consortium consisted of four groups: one at the Sanger Institute, led by Richard Durbin; one at Cold Spring Harbor laboratory, led by Lincoln Stein; one at Washington University St. Louis, led by John Speith; and one at the California Institute of Technology, led by Paul Sternberg (lead principle investigator for the project as a whole).
WormBase and parasite genomics
In 2010, Richard Durbin was appointed as joint head of human genetics at the Sanger Institute. In response to this, and consistent with a general shift in research interests over the last several years, Dr. Durbin took the decision to step down from the WormBase consortium. He retains a strong connection to the project in an advisory capacity.
The WTSI continues to participate in WormBase via new consortium member Matthew Berriman. Dr. Berriman’s research programme into parasite genomics and Neglected Tropical Diseases uses C. elegans as a model for the development of effective methodologies for the genome sequencing of parasitic worms. His involvement in WormBase aligns with the one of the key strategic goals of the project: to provide a resource that is useful and accessible to scientists working on non-Caenorhabditis nematodes.
Related groups
Data use
This sequencing centre plans on publishing the completed and annotated sequences in a peer-reviewed journal as soon as possible. Permission of the principal investigator should be obtained before publishing analyses of the sequence/open reading frames/genes on a chromosome or genome scale. See our data sharing policy.