Molecular Cytogenetics
Scientific Operations
Archive Page
This page is maintained as a historical record and is no longer being updated.
 Fluorescence in situ hybridisation (FISH) is a laboratory technique for detecting and locating specific DNA sequences on metaphase chromosomes and interphase nuclei. The technique relies on exposing the chromosome to a small DNA sequence called a probe that has a fluorescent molecule attached to it. The probe will only anneal to the complementary sequences under optimised condition. Simultaneous hybrisation of multiple probes, each labelled with a different fluorescent dye or a different combination of multiple dyes, allows multiple targets in a single specimen to be visualised in one hybridisation. FISH thus enables the link between genomics and cyotgeneitcs.
Fluorescence in situ hybridisation (FISH) is a laboratory technique for detecting and locating specific DNA sequences on metaphase chromosomes and interphase nuclei. The technique relies on exposing the chromosome to a small DNA sequence called a probe that has a fluorescent molecule attached to it. The probe will only anneal to the complementary sequences under optimised condition. Simultaneous hybrisation of multiple probes, each labelled with a different fluorescent dye or a different combination of multiple dyes, allows multiple targets in a single specimen to be visualised in one hybridisation. FISH thus enables the link between genomics and cyotgeneitcs. Nearly all our services rely on FISH, including physical mapping by metaphase-, interphase- and fibre-FISH; karyotyping of cancer and reprogrammed cell lines by Multiplex-FISH, studying the evolutionary chromosome rearrangement by cross-species chromosome painting etc.
Currently we can provide the following FISH services:
1. Physical assigment of BACs, fosmids, transgenes, cDNA clones onto metaphase chromosomes.
2. High-resolution mapping and gap sizing by fibre-FISH with single-molecule DNA fibres and extended chromomatin fibres.
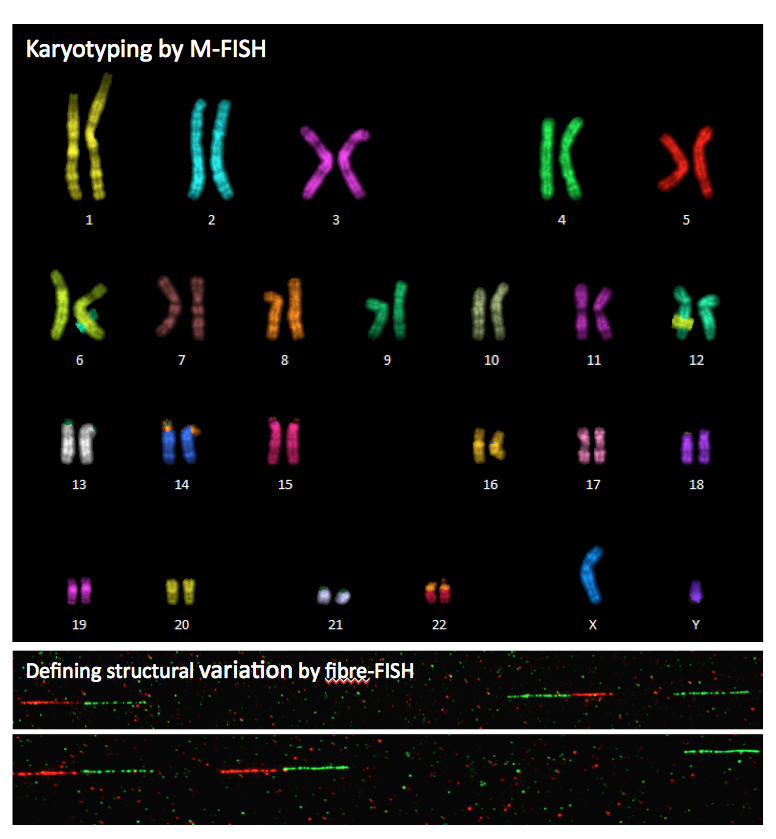
3. Karyotyping by combined M-FISH and inverted DAPI-banding.
4. Validation of structural and copy number variations.
5. Multi-directional chromosome painting
Please submit your requests via: fish-requests@sanger.ac.uk
Previous core team members

Ms Beiyuan Fu
Senior Research Assistant

Sandra Louzada Gomes Pereira
Senior Research Assistant
