Alumni
This person is a member of Sanger Institute Alumni.
Emma Carpenter joined the Sanger Institute in 2017 as an Advanced Research Assistant in Marcus Lee’s Group, before joining the Wellcome PhD programme at the Sanger Institute in 2018. She completed three rotations with Gavin Wright, Dominic Kwiatkowski, and Mathew Garnett before re-joining Marcus Lee’s group. Emma defended her PhD thesis “The influence of genetic background on drug resistance in the malaria parasite Plasmodium falciparum” in July 2023, and begins a new job at Agenus in August 2023.
My PhD project involved dissecting out minor genomic variants that contribute to, and modify, antimalarial drug resistance in the malaria parasite Plasmodium falciparum. I performed over 1,000 dose-response assays using liquid handling robots to generate confident, statistically robust conclusions, and genetically edited over 30 P. falciparum cell lines using CRISPR/Cas9. I performed a high-throughput screen of edited parasites, utilising Next-Generation Sequencing to deconvolute individual parasite growth, thereby allowing linkage analysis to be performed at scale in this organism for the first time.
–
During my time as an Advanced Research Assistant, I worked with colleagues in 8 different Universities and Institutions as part of the Malaria Drug Accelerator (MalDA) Consortium, consisting of the University of California San Diego, Columbia University, The Pennsylvania State University, Washington University in St. Louis, Harvard School of Public Health, Wellcome Sanger Institute, GlaxoSmithKline, The Medicines for Malaria Venture, and the Bill & Melinda Gates Foundation, which funded our project. Our aim is to streamline the process of:
- identifying anti-malarial compounds
- determining their mode of action
- developing promising compounds into new antimalarial drugs.
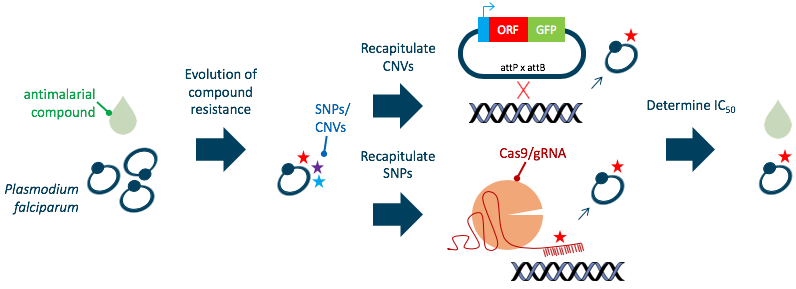
Ongoing work has identified mutations in Plasmodium falciparum, the malaria parasite, that are associated with resistance to a panel of chemical compounds. I recapitulated single alleles into a reference P. falciparum strain to determine the precise contribution of candidate mutations to the resistance phenotype: I participated in the entire validation workflow; design and generation of CRISPR-Cas9/knock-in integration plasmids, transfection into parasites, genotyping of the resulting lines, and phenotyping by dose-response assays. In a year and a half, I generated 67 plasmids, transfected 92 times, and performed over 100 drug assays.