Gene study pinpoints superbug link between people and animals
Scientists have shed light on how a major cause of human and animal disease can jump between species, by studying its genes.
The findings, published today (23 July) in Nature Ecology & Evolution, reveal fresh insights into how new disease-causing strains of the bacteria – called Staphylococcus aureus – emerge.
Experts say the research could help improve the use of antibiotics and design better strategies for limiting the spread of disease.
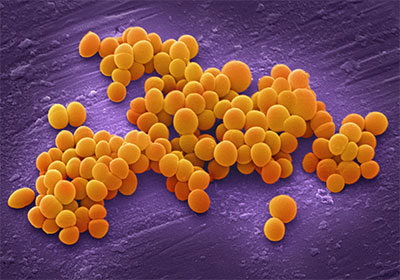
Antibiotic-resistant strains of the bacteria, such as MRSA, are a major cause of hospital-acquired infections.
The bacteria is also a major burden for the agricultural industry as it causes diseases such as mastitis in cows and skeletal infections in broiler chickens.
A team led by the University of Edinburgh’s Roslin Institute, along with their collaborators at the Wellcome Sanger Institute and elsewhere, analysed the entire genetic make-up of more than 800 strains of S. aureus that were isolated from people and animals.
The researchers sought to investigate the evolutionary history of the bacteria and key events that had allowed it to jump between species.
They found that humans were the likely original host for the bacteria. The first strains capable of infecting livestock emerged around the time animals were first domesticated for farming.
Cows have been a source of strains that now cause infections in human populations worldwide, the study found. The researchers say this highlights the importance of disease surveillance in people and animals in order to spot strains that could cause major epidemics.
The analysis revealed that each time the bacteria jumps species, it acquires new genes that enable it to survive in its new host. In some cases, these genes can also confer resistance to commonly used antibiotics.
Genes linked to antibiotic resistance are unevenly distributed among strains that infect humans compared with those that infect animals, the study found. The researchers say this reflects the distinct practices linked to antibiotic usage in medicine and agriculture.
Investigating how the bacteria are affected by genetic changes that occur after it jumps species could reveal opportunities to develop new anti-bacterial therapies, the researchers say.
It could also help to inform better strategies for managing infections to reduce the risk of transmission to people, and slow the emergence of antibiotic resistance.
“This study shows how human bacteria can jump to animals and back, and reinforces that genomic surveillance is critical in identifying new, emerging strains of drug-resistant bacteria. This information could help with designing more effective ways to prevent bacterial transfer in farms and better antibiotic practices.”
Dr Julian Parkhill An author of the study from the Wellcome Sanger Institute
“This study has been a real collaborative effort between numerous research groups in the UK and beyond. Our findings provide a framework to understand how some bacteria can cause disease in both humans and animals and could ultimately reveal novel therapeutic targets.”
Professor Ross Fitzgerald Group Leader at the University of Edinburgh’s Roslin Institute and Director of Edinburgh Infectious Disease
More information
Publication:
Emily Richardson et al. (2018) Gene exchange drives the ecological success of a multi-host bacterial pathogen. Nature Ecology & Evolution. DOI: 10.1038/s41559-018-0617-0
Funding:
The study was supported by BBSRC, MRC and Wellcome.
The Roslin Institute receives strategic funding from the Biotechnology and Biological Sciences Research Council.
Selected websites
The Wellcome Sanger Institute
The Wellcome Sanger Institute is one of the world's leading genome centres. Through its ability to conduct research at scale, it is able to engage in bold and long-term exploratory projects that are designed to influence and empower medical science globally. Institute research findings, generated through its own research programmes and through its leading role in international consortia, are being used to develop new diagnostics and treatments for human disease. To celebrate its 25th year in 2018, the Institute is sequencing 25 new genomes of species in the UK. Find out more at www.sanger.ac.uk or follow @sangerinstitute
Wellcome
Wellcome exists to improve health for everyone by helping great ideas to thrive. We’re a global charitable foundation, both politically and financially independent. We support scientists and researchers, take on big problems, fuel imaginations and spark debate. wellcome.org


