Olorin
Olorin is an interactive filtering tool for next generation sequencing data coming from the study of large complex disease pedigrees.
About
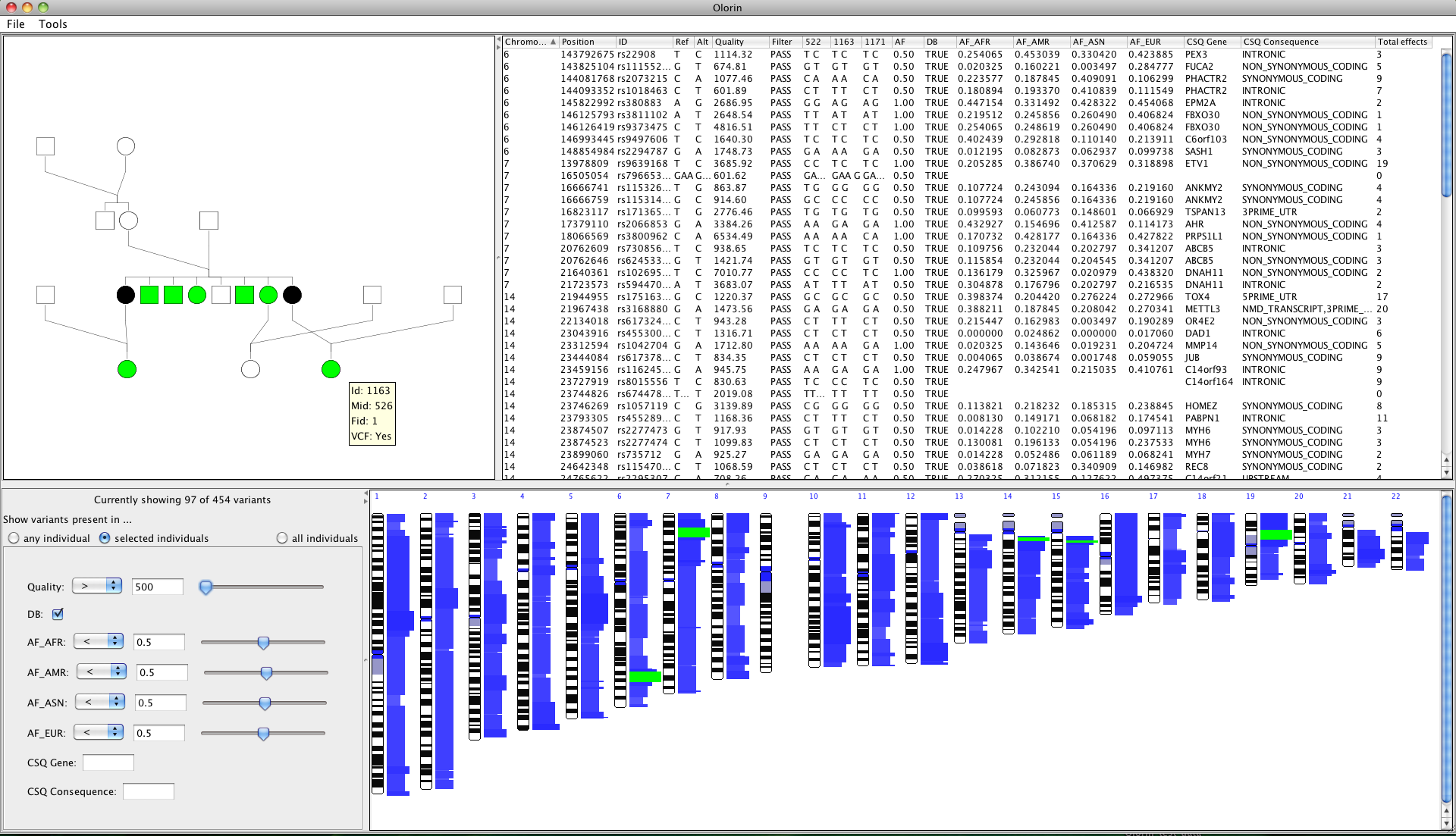
Olorin display
Downloads
System requirements
Olorin is written in Java and will work on any platform with Java 1.6 or later.
- The project is open source and is being hosted on Sourceforge
- Olorin application
- Olorin source code
- Olorin documentation
Further information
Loading data
-
Why can Olorin not find my VCF index?
Where a VCF file is named in the following way: example.vcf.gz
Olorin expects the VCF index file to be named: example.vcf.gz.tbi
What should I do if Olorin has problems fetching variants from the VCF?
First test that your VCF file was successfully indexed, you can do this on the command line using tabix and the following command:
-
tabix example.vcf.gz 1:0-1000000
This command should print out all the variants in the first megabase of chromosome one in your VCF. Try this command on a region of the genome that you know contains variants, if this does not work then try indexing your VCF again. If this does not help then you should also check that your VCF is in a valid format. You can check if your VCF is valid using VCFtools.
-
What options does MERLIN require to produce output for Olorin?
- Olorin requires the .flow files to be in the horizontal format for outputting haplotypes, MERLIN should therefore be run using the –horizontal flag.
Display
-
How can I make the genome wide ideogram plot larger?
- You can use the clickable expanding arrows found in each of the panel dividers allow each of the panels to be fully expanded to the full X or Y dimension of the Olorin window.
-
How can I identify the sample name of a node in the pedigree?
Holding the pointer over the point of interest will show which sample corresponds to that node.
Other
-
What do I do if Olorin runs out of memory?
If Olorin needs more memory to load a data set then users can launch the program with additional memory from the command line by issuing the following command (increasing the -Xmx value as necessary):
java -Xmx1024m -jar Olorin.jar
Contact
If you need help or have any queries, please contact us using the details below.
For questions or comments, please contact James Morris or Jeff Barrett.
Sanger Institute Contributors
Previous contributors

James Morris
Research Assistant