
Dr Vladimir Kiselev
Cellular Genetics Informatics Team Leader
Alumni
This person is a member of Sanger Institute Alumni.
Our team provides efficient access to cutting-edge analysis methods, environments and pipelines for Cellular Genetics programme, which leads and is involved in a number of world-class research initiatives including the Human Cell Atlas, Human Induced Pluripotent Stem Cell Initiative and Human BioMolecular Atlas Program. We work with large amounts of single-cell RNA sequencing and microscopy imaging data.
My timeline
Senior Staff Scientist, Sanger Institute
Staff Scientist, Sanger Institute
PostDoc in statistical method development for scRNA-Seq data, Sanger Institute
PostDoc in Systems Biology and Bioinformatics, European Bioinformatics Institute & Babraham Institute
PhD in Cell Biology, University of Edinburgh
MS in Applied Physics and Mathematics, Moscow Institute of Physics and Technology
BS in Applied Physics and Mathematics, Moscow Institute of Physics and Technology
Physico-Technical Lyceum 1, Saratov, Russia
Gymnasium 4, Saratov, Russia
Back in the USSR
My publications

Distribution of negatively charged lipids around positively charged peptide on the cell membrane

Transcriptional feedback loop in PIP3 signalling pathway via PRDM1
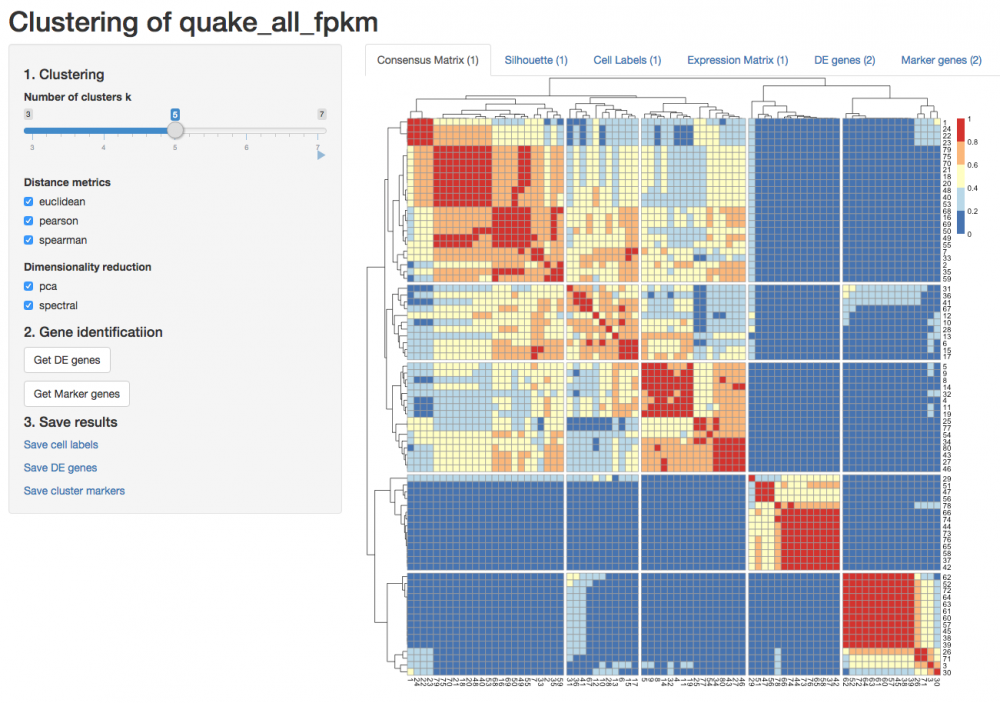
SC3 - Single-Cell Consensus Clustering tool
