Bugs as drugs
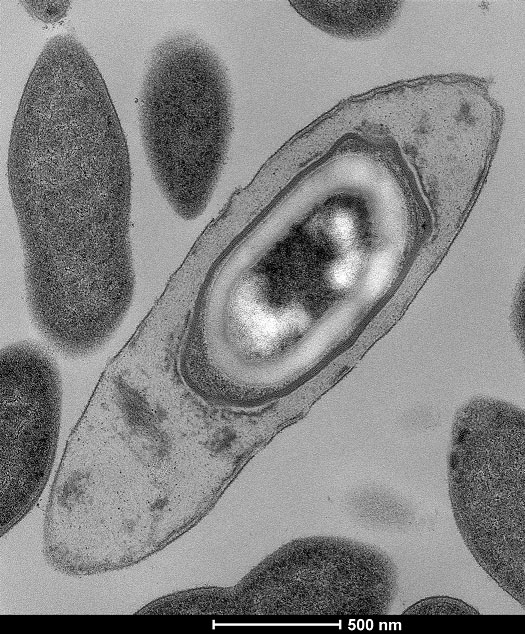
Scientists at the Wellcome Trust Sanger Institute have grown and catalogued more than 130 bacteria from the human intestine according to a study published in Nature today (Wednesday 4th May).
The researchers have developed a process to grow the majority of bacteria from the gut, which will enable scientists to understand how our bacterial ‘microbiome’ helps keep us healthy. Imbalances in our gut microbiome can contribute to complex conditions and diseases such as obesity, Inflammatory Bowel Disease, Irritable Bowel Syndrome and allergies. This research will allow scientists to start to create tailor-made treatments with specific beneficial bacteria.
Research in this field has expanded greatly in recent years with the intestinal microbiome being termed a ‘forgotten organ’, such is its importance to human health. Approximately 2 per cent of a person’s body weight is due to bacteria. Many of these bacteria are sensitive to oxygen and are difficult to culture in the laboratory, so until now it has been extremely difficult to isolate and study them.
“It has become increasingly evident that microbial communities play a large role in human health and disease. By developing a new process to isolate gastrointestinal bacteria, we were able to sequence their genomes to understand more about their biology. We can also store them for long periods of time making them available for further research.”
Hilary Browne Based in the Host-Microbiota Interactions Laboratory, at the Wellcome Trust Sanger Institute
Antibiotics wipe out our gut bacteria – killing both the pathogen targets and the beneficial bacteria too. There is then the potential for less desirable bacteria, such as those with antibiotic resistance, to repopulate the gut faster than the beneficial bacteria, leading to further health issues, such as Clostridium difficile infection.
Current treatment for C. difficile infection can involve transplants of faeces from healthy people, to repopulate the gut. However this treatment is far from ideal. Using the library of new bacteria, Dr Trevor Lawley and his team at the Sanger Institute are hoping to create a pill, containing a rationally selected, defined mix of bacteria, which could be taken by patients and replace faecal transplants.
“The extensive database of genomes we have generated from these bacteria is also essential for studying which bacteria are present or absent in people with gastrointestinal conditions. Now we can start to design mixtures of therapeutics candidates for use in these diseases.”
Dr Sam Forster from the Sanger Institute and Hudson Institute of Medical Research in Australia
For the first time The researchers also looked at the proportion of bacteria that form spores within the gut. Spores are a form of bacterial hibernation allowing some bacteria to remain dormant for long periods of time. They found approximately one third of the gut microbiota from a healthy person produced spores that allow bacteria to survive in the open air and potentially move between people. This provides a means of microbiota transmission that has not been considered before and could imply that health and certain diseases could be passed, not just through human genetics, but also via the microbiome.
“Being able to cast light on this microbial ‘Dark matter’ has implications for the whole of biology and how we consider health. We will be able to isolate the microbes from people with a specific disease, such as infection, cancers or autoimmune diseases, and study these microbes in a mouse model to see what happens. Studying our ‘second’ genome, that of the microbiota, will lead to a huge increase in our understanding of basic biology and the relationship between our gut bacteria and health and disease.”
Dr Trevor Lawley, group leader at the Sanger Institute
More information
Related publication
Samuel C. Forster et al. (2016). HPMCD: the database of human microbial communities from metagenomic datasets and microbial reference genomes. Nucleic Acids Research. DOI: 10.1093/nar/gkv1216
Funding
This work was supported by the Wellcome Trust [098051]; the United Kingdom Medical Research Council [PF451] and the Australian National Health and Medical Research Council [1091097].
Participating Centres
- Host-Microbiota Interactions Laboratory, Wellcome Trust Sanger Institute, Hinxton, UK,
- Centre for Innate Immunity and Infectious Diseases, Hudson Institute of Medical Research, Clayton, Victoria, Australia,
- Department of Molecular and Translational Sciences, Monash University, Clayton, Victoria, Australia
- Microbial Pathogenesis Laboratory, Wellcome Trust Sanger Institute, Hinxton, UK
Publications:
Selected websites
Hudson Institute of Medical Research
Hudson Institute of Medical Research is one of Australia’s top medical research institutes, specialising in driving innovative, cutting-edge research towards improved prevention, diagnosis and treatments for today’s greatest health challenges. The Institute is an NHMRC accredited, not-for-profit, independent medical research institute, an affiliate of Monash University and Monash Health, and a partner in the Monash Health Translation Precinct (MHTP). Hudson Institute is world-renowned for its research into reproductive and baby health and is a leading centre for research into infection and innate immunity, now closely tied to the latest new developments for cancer treatment.
The Wellcome Trust Sanger Institute
The Wellcome Trust Sanger Institute is one of the world’s leading genome centres. Through its ability to conduct research at scale, it is able to engage in bold and long-term exploratory projects that are designed to influence and empower medical science globally. Institute research findings, generated through its own research programmes and through its leading role in international consortia, are being used to develop new diagnostics and treatments for human disease. www.sanger.ac.uk
The Wellcome Trust
The Wellcome Trust is a global charitable foundation dedicated to achieving extraordinary improvements in human and animal health. We support the brightest minds in biomedical research and the medical humanities. Our breadth of support includes public engagement, education and the application of research to improve health. We are independent of both political and commercial interests.


