Almost 2,000 unknown bacteria discovered in the human gut
Researchers at the Wellcome Sanger Institute and EMBL’s European Bioinformatics Institute have identified almost 2,000 bacterial species living in the human gut. These species are yet to be cultured in the lab. The team used a range of computational methods to analyse samples from individuals worldwide.
The results, published in the journal Nature, highlight that although researchers are possibly getting closer to creating a comprehensive list of the commonly found microbes in the North American and European gut, there is a significant lack of data from other regions of the world.
The human gut is home to many species of microbes, collectively referred to as the gut microbiota. Despite extensive studies in the field, researchers are still working on identifying the individual microbial species that live in our guts and understanding what roles they play in human health.
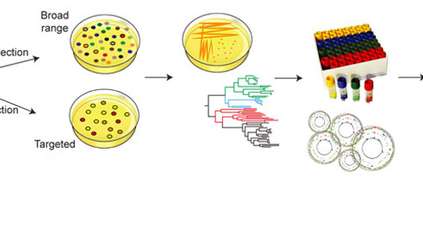
There are many reasons why some microbial species that are part of the gut microbiota have remained unknown for so long, such as a low abundance in the gut or an inability to survive outside it. By using computational methods, researchers were able to reconstruct the genomes of these bacteria.
“Computational methods allow us to understand bacteria that we cannot yet culture in the lab. Using metagenomics to reconstruct bacterial genomes is a bit like reconstructing hundreds of puzzles after mixing all the pieces together, without knowing what the final image is meant to look like, and after completely removing a few pieces from the mix just to make it that bit harder. Researchers are now at a stage where they can use a range of computational tools to complement and sometimes guide lab work, in order to uncover new insights into the human gut.”
Rob Finn Group Leader at EMBL-EBI
The research highlighted how the composition of gut bacteria differs around the world, and how important it is for the samples that we study to reflect this diversity.
“We are seeing a lot of the same bacterial species crop up in the data from European and North American populations. However, the few South American and African datasets we had access to for this study revealed significant diversity not present in the former populations. This suggests that collecting data from underrepresented populations is essential if we want to achieve a truly comprehensive picture of the composition of the human gut.”
Rob Finn Group Leader at EMBL-EBI
“Computational methods allow us to get an idea of the many bacterial species that live in the human gut, how they evolved and what kind of roles they may play within their microbial community. In this study, we leveraged the most comprehensive public databases of gastrointestinal bacteria to identify bacterial species that have not been seen before. The analysis methods we used are highly reproducible and can be applied to larger, more diverse datasets in the future, enabling further discovery.”
Alexandre Almeida Postdoctoral Fellow at EMBL-EBI and the Wellcome Sanger Institute
“Research such as this is helping us create a so-called blueprint of the human gut, which in the future could help us understand human health and disease better and could even guide diagnosis and treatment of gastrointestinal diseases.”
Trevor Lawley Group Leader at the Wellcome Sanger Institute
More information
Almeida A et al. (2019). A new genomic blueprint of the human gut microbiota. Nature. Published online: 11/02/19; DOI: 10.1038/s41586-019-0965-1
Selected websites
EMBL’s European Bioinformatics Institute (EMBL-EBI)
EMBL’s European Bioinformatics Institute (EMBL-EBI) is a global leader in the storage, analysis and dissemination of large biological datasets. EMBL-EBI helps scientists realise the potential of ‘big data’ by enhancing their ability to exploit complex information to make discoveries that benefit humankind.
EMBL-EBI is at the forefront of computational biology research, with work spanning sequence analysis methods, multi-dimensional statistical analysis and data-driven biological discovery, from plant biology to mammalian development and disease.
EMBL-EBI is part of the European Molecular Biology Laboratory (EMBL), an international, innovative and interdisciplinary research organisation funded by 25 member states and two associate member states, and are located on the Wellcome Genome Campus, one of the world’s largest concentrations of scientific and technical expertise in genomics. www.ebi.ac.uk
Wellcome Sanger Institute
The Wellcome Sanger Institute is one of the world’s leading genome centres.
Through its ability to conduct research at scale, it is able to engage in bold and long-term exploratory projects that are designed to influence and empower medical science globally. Institute research findings, generated through its own research programmes and through its leading role in international consortia, are being used to develop new diagnostics and treatments for human disease.
Find out more at www.sanger.ac.uk or follow @sangerinstitute on Twitter, Facebook, LinkedIn and on our Blog.
About Wellcome
Wellcome exists to improve health by helping great ideas to thrive.
We support researchers, we take on big health challenges, we campaign for better science, and we help everyone get involved with science and health research.
We are a politically and financially independent foundation. wellcome.org