Exomiser
The Exomiser is a Java program that finds potential disease-causing variants from whole-exome or whole-genome sequencing data.
Archive Page
This page is maintained as a historical record and is no longer being updated.
Exomiser is no longer hosted at the Sanger Institute. Data can now be found at http://data.monarchinitiative.org/exomiser/. Source code and binary distribution can be found at https://github.com/exomiser/Exomiser
This page is no longer being updated and is being maintained as a historical record of the work that was carried out the Institute
About
The functional annotation of variants is handled by Jannovar and uses UCSC KnownGene transcript definitions and hg19 genomic coordinates.
Variants are prioritised according to user-defined criteria on variant frequency, pathogenicity, quality, inheritance pattern, and model organism phenotype data. Predicted pathogenicity data is extracted from the dbNSFP resource. Variant frequency data is taken from the 1000 Genomes, ESP and ExAC datasets. Subsets of these frequency and pathogenicity data can be defined to further tune the analysis. Cross-species phenotype comparisons come from our PhenoDigm tool powered by the OWLTools OWLSim algorithm.
The Exomiser was developed by the Computational Biology and Bioinformatics group at the Institute for Medical Genetics and Human Genetics of the Charité – Universitätsmedizin Berlin, the Mouse Informatics Group at the Sanger Institute and other members of the Monarch initiative.
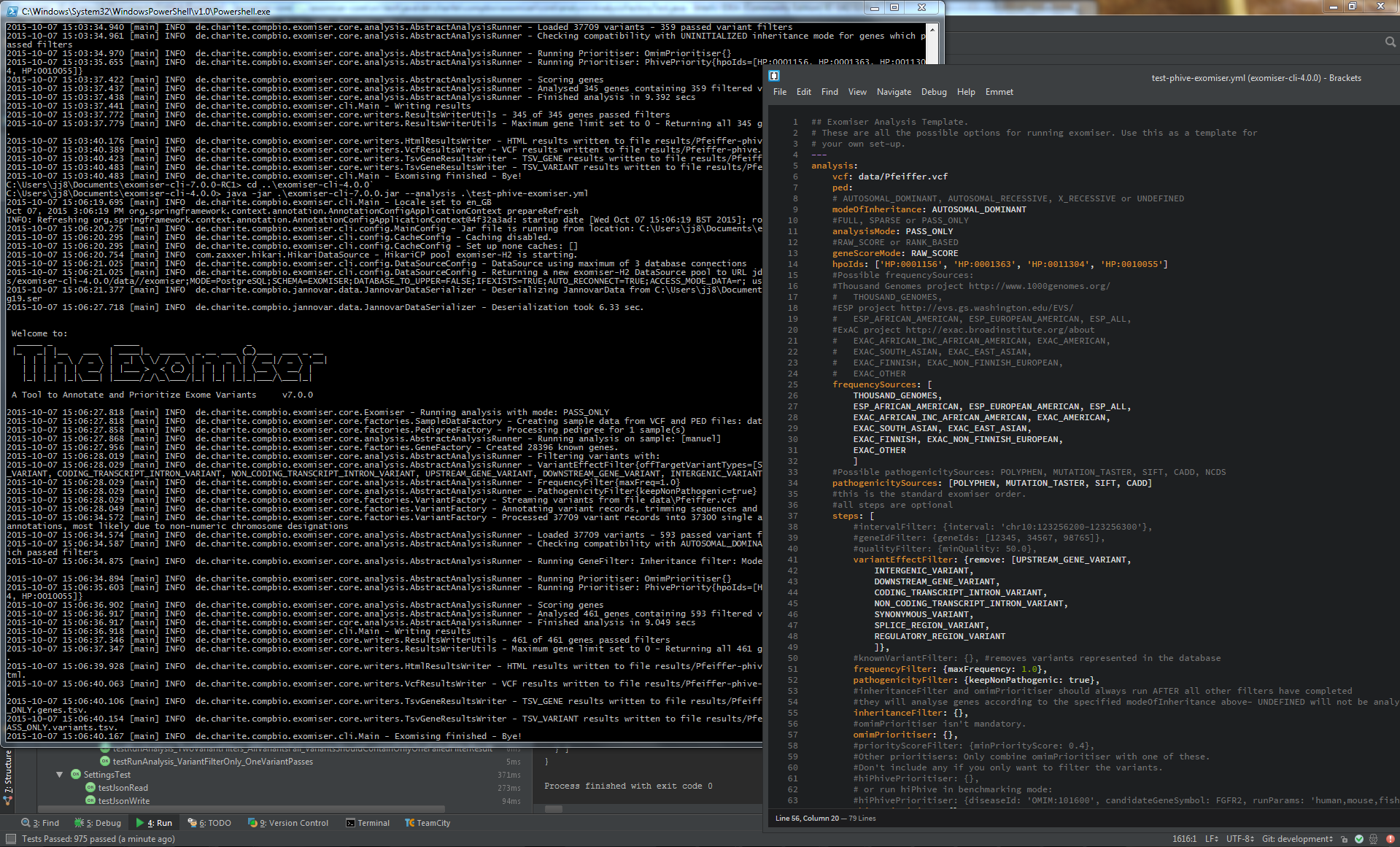
Screenshot of the Exomiser running on a test dataset from the command-line along with the accompanying YAML configuration file used for the analysis.
Downloads
Exomiser is no longer hosted at the Sanger Institute. Data can now be found at http://data.monarchinitiative.org/exomiser/. Source code and binary distribution can be found at https://github.com/exomiser/Exomiser.
Further information
Please refer to the README file on the github site for instructions on running the Exomiser.
Published under the GNU Affero General Public License, version 3
Please cite the Exomiser using:
Improved exome prioritization of disease genes through cross-species phenotype comparison.
Robinson PN, Köhler S, Oellrich A, Sanger Mouse Genetics Project, Wang K, Mungall CJ, Lewis SE, Washington N, Bauer S, Seelow D, Krawitz P, Gilissen C, Haendel M and Smedley D
Genome research 2014;24;2;340-8
PUBMED: 24162188; PMC: 3912424; DOI: 10.1101/gr.160325.113
Contact
If you need help or have any queries, please contact us using the details below.
The authors (Damian Smedley or Jules Jacobsen) are no longer at the Sanger Institute and can be contacted on the Exomiser development pages hosted on GitHub:
Sanger Institute Contributors
Previous contributors

Dr Jules Jacobsen
Senior Software Developer

Dr Damian Smedley
Former Senior Scientific Manager