Drug2Cell
Archive Page
This page is maintained as a historical record and is no longer being updated.
Technology
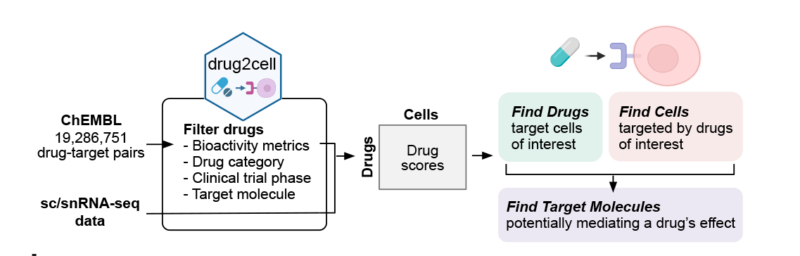
The tool takes pairs of drugs and their target molecules from the EMBL-EBI ChEMBL database leveraging single-cell profiles and the 19 million drug-target interactions. Drugs and target molecules can be queried and filtered based on quantitative bioactivity metrics, drug categories (ATC classification), clinical trial phase, and classes of molecular targets. The scores of the selected drugs in single cells or cell populations are calculated based on the target gene expressions in sn/scRNAseq data.
Once drug scores are calculated, the tool allows to:
- Find cells which are targeted by drugs of interest;
- Find drugs which target cells of interest;
- Find target molecules expressed in the target cells to infer a drug’s effect.
Advantages
Drug2cell can identify specific cellular targets of bioactive molecules with drug properties based on single-cell RNA-seq data. This offers a new perspective on drug discovery process potentially revealing hidden mechanisms of action and predicting the impact of medicines on specific cell types.
Detail
Applications/Context
- Drug2cell can enable and improve how new treatments are evaluated and how they affect organs and tissues.
- It allows for checking specific cell targets of drugs in organs of the body where single cell data is avaiable.
- In the context of the heart, the analysis identified pacemaker cell-expressed targets for non-cardiac medications with documented chronotropic effects. These included the anti-diabetic medication Liraglutide (GLP-1 analogue) and the anti-epileptic medication Perampanel (glutamate AMPA receptor inhibitor). Both drugs are known to alter heart rate, but since their targets were not known to be present in pacemaker cells prior to this study, alternative sites of action (the ANS and CNS, respectively) had been previously proposed as mediators of the side effects.
- Applying drug2cell to whole body cell atlas datasets will improve in silico screening by predicting adverse effects in cell types across all organs.
Comparable Technologies
Several single-cell studies and methods have been published which use drug-response transcriptional signatures, obtained from cell line experiments, and data-mining to predict drug effects. However, these require the development of cell type-specific in vitro models, which can fail to fully recapitulate the profiles of their in vivo human counterparts. Drug2cell is leveraging single-cell profiles from tissue/organ donors providing more physiological predictions.
Background
There is a major need to identify new therapeutic targets. Additionally, off-target activity is a major reason for drug development failure and withdrawal. Single-cell technologies such as single-cell RNA sequencing (scRNA-seq) together with associated computational tools and the growing availability of public data to resources, have the opportunity to transform drug discovery and development. New opportunities are emerging in target identification due to improved disease understanding through cell subtyping, and multiplexed functional genomics screens incorporating scRNA-seq that are enhancing target prioritization.
Drug2cell can identify specific cellular targets of bioactive molecules with drug properties based on single-cell RNA-seq data, potentially revealing hidden mechanisms of action and predicting the impact of medicines on specific cell types. We leveraged our human heart data to map drugs to target-expressing cells. Several single-cell studies and methods have been published which use drug-response transcriptional signatures, obtained from cell line experiments, and data-mining to predict drug effects. However, these require the development of cell type-specific in vitro models, which (despite their increasing complexity) can fail to fully recapitulate the profiles of their in vivo counterparts.
Using our sc/ snRNAseq data as a reference, we took pairs of drug and target molecules from the ChEMBL database (https://www.ebi.ac.uk/chembl/) to comprehensively evaluate drug target expression in user-provided single-cell data and annotated cell populations. We developed a Python package, drug2cell, to streamline the workflow and apply selected drug-target pairs to single-cell datasets. Drugs and target molecules can be filtered based on quantitative bioactivity metrics, drug categories and clinical trial phases, and target molecule classes.
Cell atlasing efforts, especially those profiling cell types at a whole-body scale, provide valuable resources for understanding the similarity and difference of drug responses across organs. These datasets surveyed cross-tissue immune (Domínguez Conde et al. 2022) and vascular (Yang et al., unpublished) cells at the adult stages, as well as a multitude of immune cell types (Suo et al. 2022) and epithelial, mesenchymal and neuronal cells (Webb et al., unpublished) at the embryonic or foetal stages. Applications of drug2cell to these atlases hold promise for charting their drug-target interaction map at scale and discovering organ-shared or specific effects for cell types of interest.
While the above datasets will be released to the public domain with respective publications, Wellcome Sanger Institute might offer early data access for commercial projects with clear societal impact.
Intellectual property
The Software is available for non-commercial use only. The Wellcome Sanger Institute is offering non-exclusive licenses to the software for comercial use.
