DNAPlotter
An interactive Java application for generating circular and linear representations of genomes.
Archive Page
This page is maintained as a historical record and is no longer being updated.
This page was archived in 2022
About
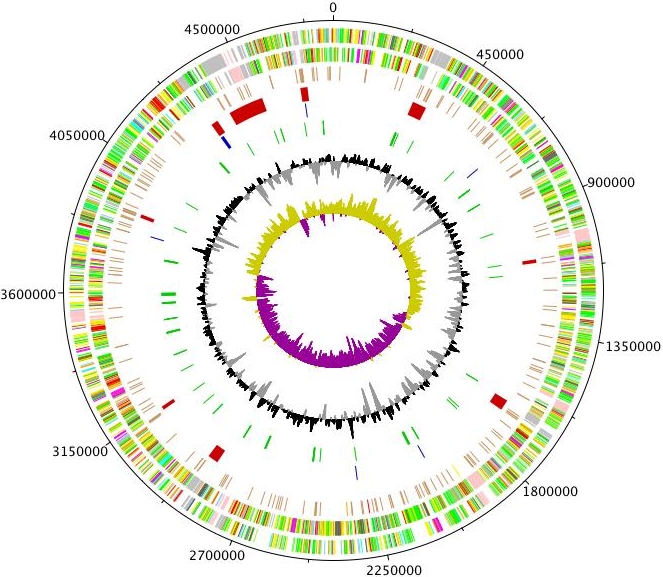
Example showing S. typhi genome
Downloads
Please see our GitHub page for download and installation instructions.
Further information
For more information and advice on using this software please see the Artemis manual and our GitHub page.
In addition, an email discussion list called artemis-users is available and posts to the list since September 2001 are archived at mail-archive.com
DNAPlotter is available under GPL3.
If you make use of this software in your research, please cite as:
DNAPlotter: circular and linear interactive genome visualization.
Carver T, Thomson N, Bleasby A, Berriman M and Parkhill J
Bioinformatics (Oxford, England) 2009;25;1;119-20
PUBMED: 18990721; PMC: 2612626; DOI: 10.1093/bioinformatics/btn578
Contact
If you need help or have any queries, please contact us using the details below.
If you need help or have any queries, please contact us using the details below.
- Email: artemis-help@sanger.ac.uk
Our normal office hours are 8:30-17:00 (GMT) Monday to Friday.