
Genome Reference Informatics
As the impact of the human reference genome assembly on biomedical research has shown, the availability of a high quality reference genome assembly is essential for the understanding of a species' biology. Our team is responsible for further improving and extending the human reference assembly, as well as the mouse and zebrafish reference assemblies. To achieve this, we are generating sequencing and mapping data from various technologies, develop software to analyse and interpret these data and apply manual curation to resolve assembly issues. We are also generating new assemblies of strains/individuals of importance for the respective community.
Current projects
- improvement of the human reference assembly, correcting errors and adding variation
- correcting and improving the draft assemblies of 16 mouse strains
- adding missing gene loci to the mouse reference assembly
- creating new zebrafish assemblies from different strains, developing techniques to enable haplotype resolution
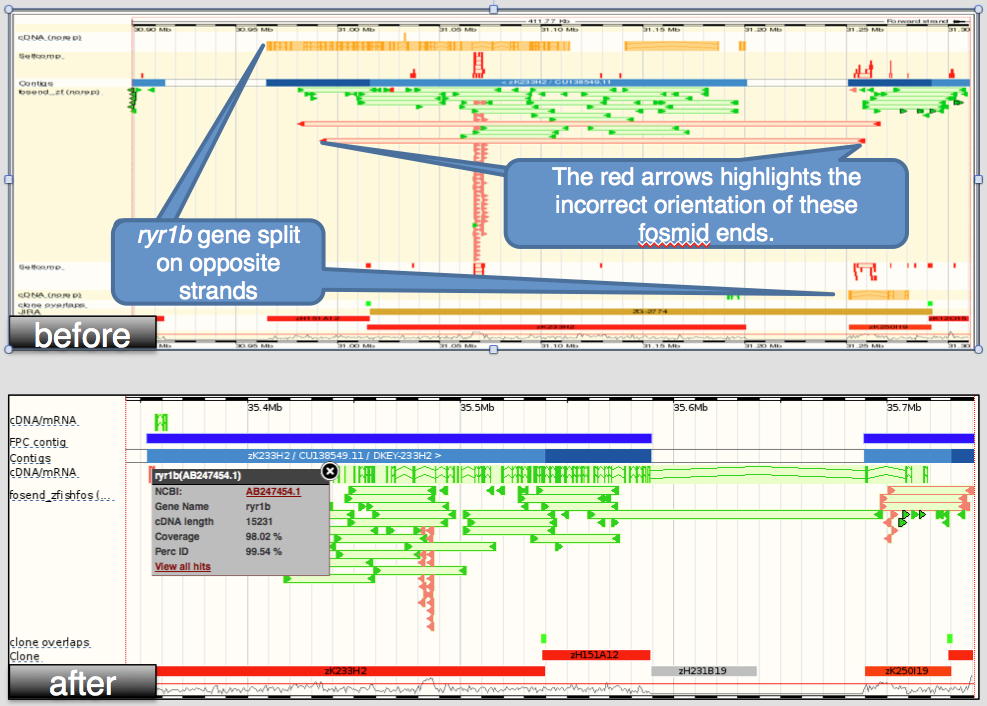
- improving the zebrafish reference assembly through addition of more clone sequence and integration with a new de novo assembly
Software
We develop software to support our research, focussing on the genome evaluation browser (gEVAL). This browser allows for an easy assessment of genome assemblies and the resolution of identified issues and is one of the main tools used by the Genome Reference Consortium, but also by e.g. the zebrafish community. We also provide Chromoview, a browser for viewing the latest assembly paths.
External partners and funders
External
Genome Reference Consortium
External