HapMap 3 is the third phase of the International HapMap project. This phase increases the number of DNA samples covered from 270 in phases I and II to 1,301 samples from a variety of human populations. This is the draft release 3.
Archive Page
This page is maintained as a historical record and is no longer being updated.
Archive Page: This page is maintained as a historical archive and is no longer being updated.
About
The definitive data are available from the HapMap ftp site. The data available from these pages at the Sanger Institute are raw unfiltered data, provided as a resource to the community.
Populations
The following population samples were studied:
- ASW – African ancestry in Southwest USA
- CEU – Utah residents with Northern and Western European ancestry from the CEPH collection
- CHB – Han Chinese in Beijing, China
- CHD – Chinese in Metropolitan Denver, Colorado
- GIH – Gujarati Indians in Houston, Texas
- JPT – Japanese in Tokyo, Japan
- LWK – Luhya in Webuye, Kenya
- MXL – Mexican ancestry in Los Angeles, California
- MKK – Maasai in Kinyawa, Kenya
- TSI – Toscani in Italia
- YRI – Yoruba in Ibadan, Nigeria
Production and AC
Genotyping
Genotyping concordance between the two platforms was 0.9931 (computed over 249889 overlapping SNPs).
Data from the two platforms was merged using PLINK (–merge-mode 1), keeping only genotype calls if there is consensus between non-missing genotype calls (that is, merged genotype is set to missing if the two platforms give different, non-missing calls).
Quality control at the individual level was performed separately for different platforms. Only individuals with QC passed genotype data on both platforms were kept in this release. The following criteria were used to keep SNPs in the data sets of this release:
Hardy-Weinberg p>0.000001 (per population) missingness <0.05 (per population) <3 Mendel errors (per population; only applies to YRI, CEU, ASW, MXL, MKK) SNP must have a rsID and map to a unique genomic location The “consensus” data set contains data for all individuals (558 males, 557 females; 924 founders and 191 non-founders), only keeping SNPs that passed QC in all populations (overall call rate is 0.998). The “consensus/polymorphic” data set has In all genotype files, alleles are expressed as being on the (+/fwd) strand of NCBI build 36.
PCR Resequencing
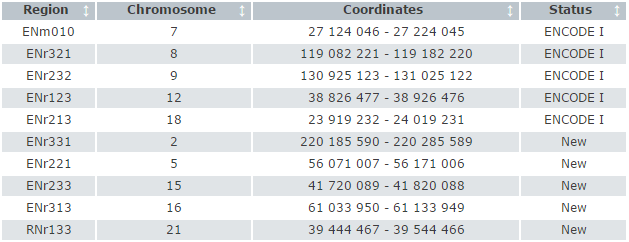
The sequence based variant calls were generated by tiling with PCR primer sets spaced approximately 800 bases apart across the regions shown in the table.
Following filtration of low quality reads the data were analyzed with SNP Detector version 3, for polymorphic site discovery and individual genotype calling. Various QC filters were then applied. Specifically, we filtered out PCR amplicons with too many SNPs, and SNPs with discordant allele calls in mutliple amplicons. We also filtered out SNPs with low completeness in samples, or with too many conflicting genotype calls in two different strands.
In the “QC+” data set, we applied the HapMap QC parameters, specifically, we filtered out samples with low completeness, and filtered out SNPs with low call rate in each population (<80%) and not in HWE (P<0.001). In the QC+ data set, overall false positive rate is ~3.2%, based on limited number of validation assays.
Caveats
Genotyping
Missing from this release are Illumina SNPs that are A/T or C/G due to strandedness issues. Missing from this release are Illumina SNPs that are mitochondrial (as they do not have rsIDs). There may be few remaining SNPs (Illumina) in this release that are still on (-/rev) strand of NCBI build 36, but they are not A/T or C/G SNPs, so easy to identify downstream.
PCR Resequencing
All variant calls have not been validated: we estimate that there is currently a false positive rate of ~12% among all calls, with a slightly higher rate (~14%) if considering just the singletons. Additional validation is ongoing PCR sequencing of additional samples (Masai) is also ongoing.
Analysis Plans
Below are the analysis plans that the consortium pursuing:
- SNP allele frequency estimation
- Population differentiation
- Linkage disequilibrium analysis
- SNP Tagging
- Imputation efficiency
- Genomic locations of human CNVs
- Genotypes for CNVs
- Population genetic properties of CNVs (allele frequencies, population differentiation, etc.)
- Mutation rate (frequency of de novo CNV) and potential mutational mechanisms
- Linkage disequilibrium properties of CNVs
- Tagging and imputation of CNVs
- Signals of selection around CNVs
- Association of SNPs and CNVs with expression phenotypes
Affiliated Sites
External
International HapMap Project
External
National Human Genome Research Institute
Co-funder of phase 3 of the International HapMap Project
Data use
This sequencing centre plans on publishing the completed and annotated sequences in a peer-reviewed journal as soon as possible. Permission of the principal investigator should be obtained before publishing analyses of the sequence/open reading frames/genes on a chromosome or genome scale. See our data sharing policy.