Large-scale study raises hopes for development of E. coli vaccine
The largest ever study of the bacterium enterotoxigenic Escherichia coli (ETEC) has raised hopes that a global vaccine can be developed. This bacterium causes 400 thousand deaths and 400 million cases of diarrhoea each year in low- and middle-income countries as well as misery to many travellers to these regions.
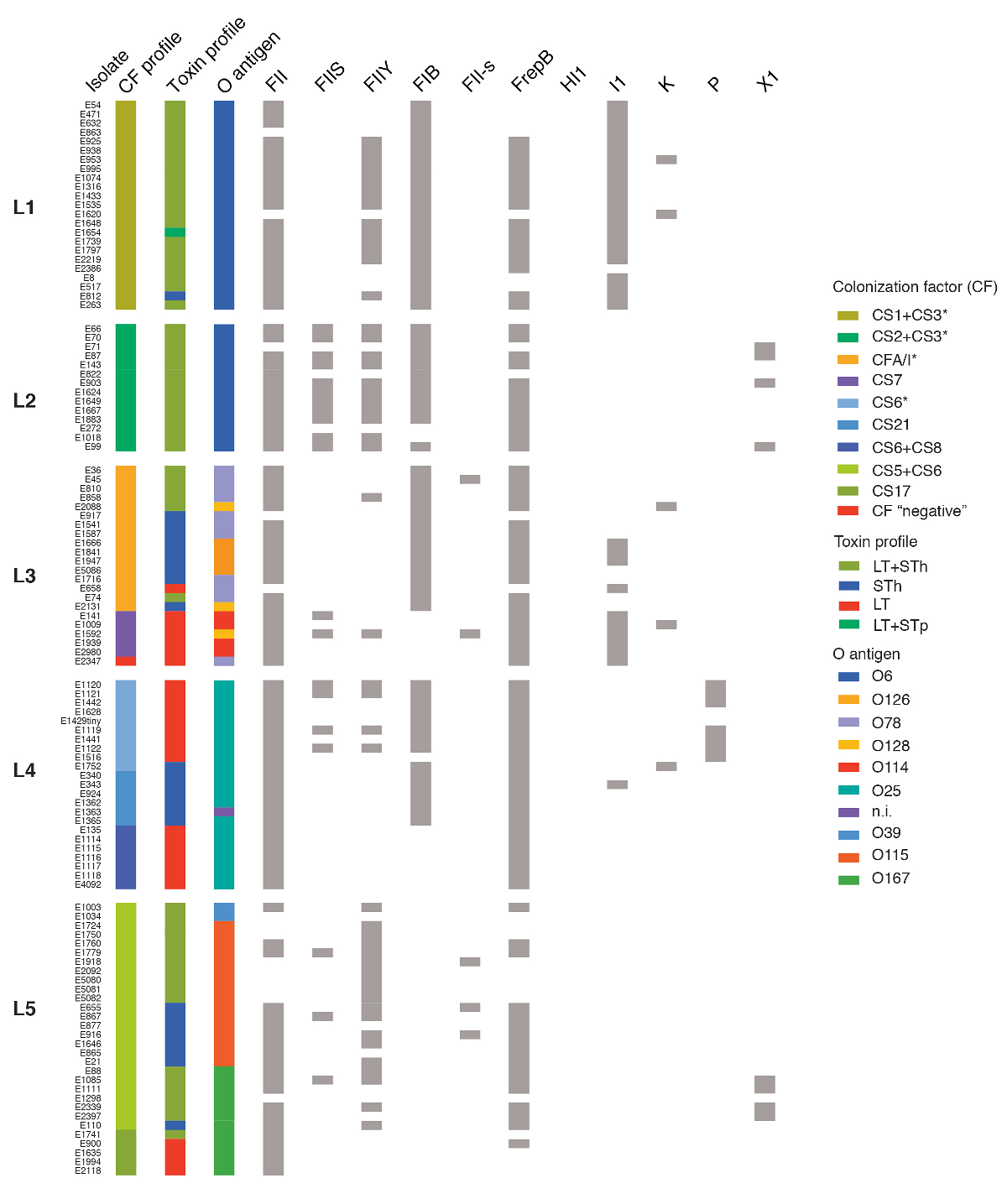
Analysis based on whole-genome sequence data for 362 different ETEC strains that span 30 years and 20 countries has overturned previous thinking about the bacterium’s diversity. By comparing the way strains attach to human stomach lining, driven in part by the colonisation factor, researchers found that strains clustered into surprisingly closely related groups. This means that the bacteria that cause disease in countries in Latin America are actually very similar to those in Africa and Asia. Some E. coli have spread globally from a common source, a previously unrecognised phenomenon.
“This research strengthens our belief that it is possible to target a broad range of ETEC groups with one vaccine. By targeting the most prevalent colonisation factors in these lineages, we stand a chance of developing a vaccine that will reduce the disease burden caused by this bacterium. This work is now underway at the Sanger Institute.”
Professor Gordon Dougan Senior author from the Wellcome Trust Sanger Institute
The close relationship between samples in Asia, Africa and the Americas suggests that a vaccine could be effective worldwide. Samples taken from children and adults in regions where the diarrhoeal disease is endemic and from people who had travelled to these areas were also similar, suggesting that a potential vaccine would work across patient groups.
Large-scale studies like this enable researchers to plot the phylogenetic structure, the pattern of global emergence and evolution, of lethal bacteria such as ETEC. While ETEC was previously thought to vary widely across the world, this phylogeny traces many of the 21 different lineages back their origins in an individual bacterium that acquired the genetic information required to infect humans and then spread, in part due to global travel. According to this data, the emergence of these lineages probably took place between 51 and 170 years ago, a finding that suggests that the lineages are stable and unlikely to become quickly resistant to vaccines.
“Our work to develop a vaccine has received a real boost from this research. This data gives us confidence that a global, future-proof solution is within our grasp.”
Dr Åsa Sjöling An author from the University of Gothenburg and Karolinska Institutet, Sweden
Detailed genetic information on each lineage has been made publicly available to the research community, enabling scientists to learn more about the bacteria and to track its future spread and evolution. Among other studies, researchers will use this data to learn more about the 130 ETEC samples that had colonisation factors that had never been identified before.
“This dataset is exceptional. Researchers around the world will be able, from these whole-genome sequences, to draw out data about the virulence profiles and other proteins that may be important for virulence and that these strains could have in common. All of this research helps us to identify the shared weaknesses in ETEC bacteria that we can exploit with vaccines.”
Astrid von Mentzer First author from the University of Gothenburg and the Sanger Institute
“Whole genome sequencing has enabled us to quantify what makes an ETEC in unprecedented detail; information that is vital in order to target the development of new vaccines. The challenge now is to distil, from the vast wealth of data locked up in this dataset, the information needed to develop these new treatments to fight this key pathogen.”
Study co-author Dr Thomas Connor from Cardiff University
More information
Funding
This work was supported by the Wellcome Trust (grant 098051), the Swedish Research Council (grants 2012-3464 and 2011-3435), the Swedish Strategic Foundation (grant SB12-0072) and the European Research Council (grant 239784).
Participating Centres
- Department of Microbiology and Immunology, Sahlgrenska Academy, University of Gothenburg, Gothenburg, Sweden;
- Wellcome Trust Sanger Institute, Hinxton, UK;
- Organisms and Environment Division, Cardiff University School of Biosciences, Cardiff University, Cardiff, UK;
- Centre of Infection Medicine, Institute of Microbiology and Epizootics, Freie Universität Berlin, Berlin, Germany;
- Interdisciplinary Research Organization, University of Miyazaki, Miyazaki, Japan;
- Department of Microbiology and Immunology, University of Maryland School of Medicine, Baltimore, Maryland, USA;
- Department of Mathematics and Statistics, University of Helsinki, Helsinki, Finland;
- Department of Microbiology, Tumor and Cell Biology, Karolinska Institutet, Stockholm, Sweden.
Publications:
Selected websites
The University of Gothenburg
The University of Gothenburg tackles society’s challenges with diverse knowledge. 37,000 students and 6,000 employees make the University a large and inspiring place to work and study, with a continuous flow of new knowledge and ideas. Strong research and attractive study programmes attract scientists and students from around the world. The University of Gothenburg is environmentally certified and works actively for sustainable development. With new knowledge and new perspectives, the University contributes to a better future.
Cardiff University
Cardiff University is recognised in independent government assessments as one of Britain’s leading teaching and research universities and is a member of the Russell Group of the UK’s most research intensive universities. Among its academic staff are two Nobel Laureates, including the winner of the 2007 Nobel Prize for Medicine, University Chancellor Professor Sir Martin Evans. Founded by Royal Charter in 1883, today the University combines impressive modern facilities and a dynamic approach to teaching and research. The University’s breadth of expertise encompasses: the College of Arts, Humanities and Social Sciences; the College of Biomedical and Life Sciences; and the College of Physical Sciences and Engineering, along with a longstanding commitment to lifelong learning. Cardiff’s four flagship Research Institutes are offering radical new approaches to cancer stem cells, catalysis, neurosciences and mental health and sustainable places.
The Wellcome Trust Sanger Institute
The Wellcome Trust Sanger Institute is one of the world’s leading genome centres. Through its ability to conduct research at scale, it is able to engage in bold and long-term exploratory projects that are designed to influence and empower medical science globally. Institute research findings, generated through its own research programmes and through its leading role in international consortia, are being used to develop new diagnostics and treatments for human disease.
The Wellcome Trust
The Wellcome Trust is a global charitable foundation dedicated to achieving extraordinary improvements in human and animal health. We support the brightest minds in biomedical research and the medical humanities. Our breadth of support includes public engagement, education and the application of research to improve health. We are independent of both political and commercial interests.


