Deciphering the life history of 21 breast cancers
In a study published online by Cell on 17 May 2012, researchers have deciphered the life history imprinted on the genome of 21 breast cancers.
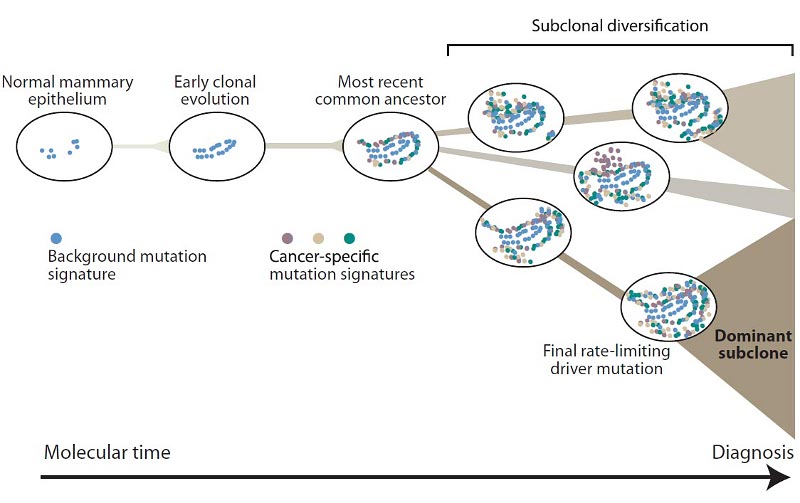
The team, led by researchers from the Wellcome trust Sanger Institute, found that mutational processes evolve across the lifespan of a breast tumour, with distinctive patterns of mutation emerging late in the process, contributing to a large amount of variation. As the breast cells accumulate many thousands of mutations, the developing cancer starts to diverge into families of genetically related cells. By the time the cancer is diagnosed, one of these families has become the dominant population in the tumour.
Each time a cell mutates, it leaves a distinctive stamp inscribed in its genome. Each mutational process leaves distinctive marks on the genome of a cancer. As these changes accumulate one on top of one another, archaeological layers of mutations build up, leaving a record that can be deciphered to allow a glimpse of the cancer’s life history.
“We wanted to see if we could decipher this history encrypted in the genome of each of the cancers. All cancers are made up of a collection of families of cells. Whole-genome sequences reveal the genetic record of their emergence over time and allow us to trace the divergence of a cell to form the different families.”
Dr Peter Campbell Head of Cancer Genetics and Genomics at the Wellcome Trust Sanger Institute
As cells divide and multiply, mutations accumulate naturally: the team found that, as these mutations accumulate in a cell, the changes drive divergence of the cancer into different subgroups or clones. The scientists showed that emergence of different clones was universal in the breast cancers they examined.
They also showed that one sub-clone becomes the dominant population of cancer cells. This dominant sub-clone accounts for more than 50 per cent of tumour cells in all of the breast cancers analysed, and it is only when this sub-clone has grown sufficiently populous that the tumour becomes clinically detectable. It differs from the other clones present in a tumour by many hundreds to thousands of mutations, indicating that this final stage in a cancer’s emergence may take quite some time.
“We undertook a deep excavation of the tumour DNA, revealing for the first time the fine structure of breast cancer genomes. We took all this data and integrated it to build an evolutionary tree of the cancers.
“This method allowed us to determine when divergence occurs, what processes are involved in the different stages of cancer evolution and the proportion of each sub-clone present in the tumours.”
Dr Serena Nik-Zainal First author on the studies, from the Wellcome Trust Sanger Institute
The team found that some mutational processes act throughout the evolution of the cancer and some processes are present only quite late in the development of the cancer once divergence has occurred.
For example, four of the cancers had many extra copies of the gene that is the target of the Herceptin drug. The team found that the first few extra copies of the gene were gained very early in the development of the breast cancer, but it took the cancer a much longer period of time than expected to accumulate all of the extra 20–30 copies.
“Current cancer treatments do not take sub-clonal diversity into account and often target only the dominant sub-clone. This leaves the possibility that one of the minor sub-clones will then replicate and become dominant, leading to re-occurrence of the tumour.
“Understanding sub-clonal diversity in breast cancer is a pivotal part of treating this destructive cancer in the most efficient way. This study forms the basis to identify sub-clones both minor and dominant.”
Professor Mike Stratton Co-author and Director of the Wellcome Trust Sanger Institute
These findings have significant implications for our understanding of how breast cancers develop over the decades prior to diagnosis. The next step for this research is to sequence more genomes and cancer types, and also to refine current methods.
More information
Funding
A full list of funding agencies can be found in the papers.
Participating Centres
A full list of participating centres can be found in the papers.
Publications:
Selected websites
Breakthrough Breast Cancer
Breakthrough Breast Cancer is dedicated to improving and saving lives through finding the causes of breast cancer, enabling early detection, ensuring precise diagnosis, discovering new and better treatments and improving medical services. Breakthrough Breast Cancer funds groundbreaking research, campaign for better services and treatments and raise awareness of the signs and symptoms of breast cancer. Through this work the charity believes passionately that breast cancer can be beaten and the fear of the disease removed for good.
The Wellcome Trust Sanger Institute
The Wellcome Trust Sanger Institute is one of the world’s leading genome centres. Through its ability to conduct research at scale, it is able to engage in bold and long-term exploratory projects that are designed to influence and empower medical science globally. Institute research findings, generated through its own research programmes and through its leading role in international consortia, are being used to develop new diagnostics and treatments for human disease.
The Wellcome Trust
The Wellcome Trust is a global charitable foundation dedicated to achieving extraordinary improvements in human and animal health. We support the brightest minds in biomedical research and the medical humanities. Our breadth of support includes public engagement, education and the application of research to improve health. We are independent of both political and commercial interests.


