National project paves way for clinical genetic diagnosis
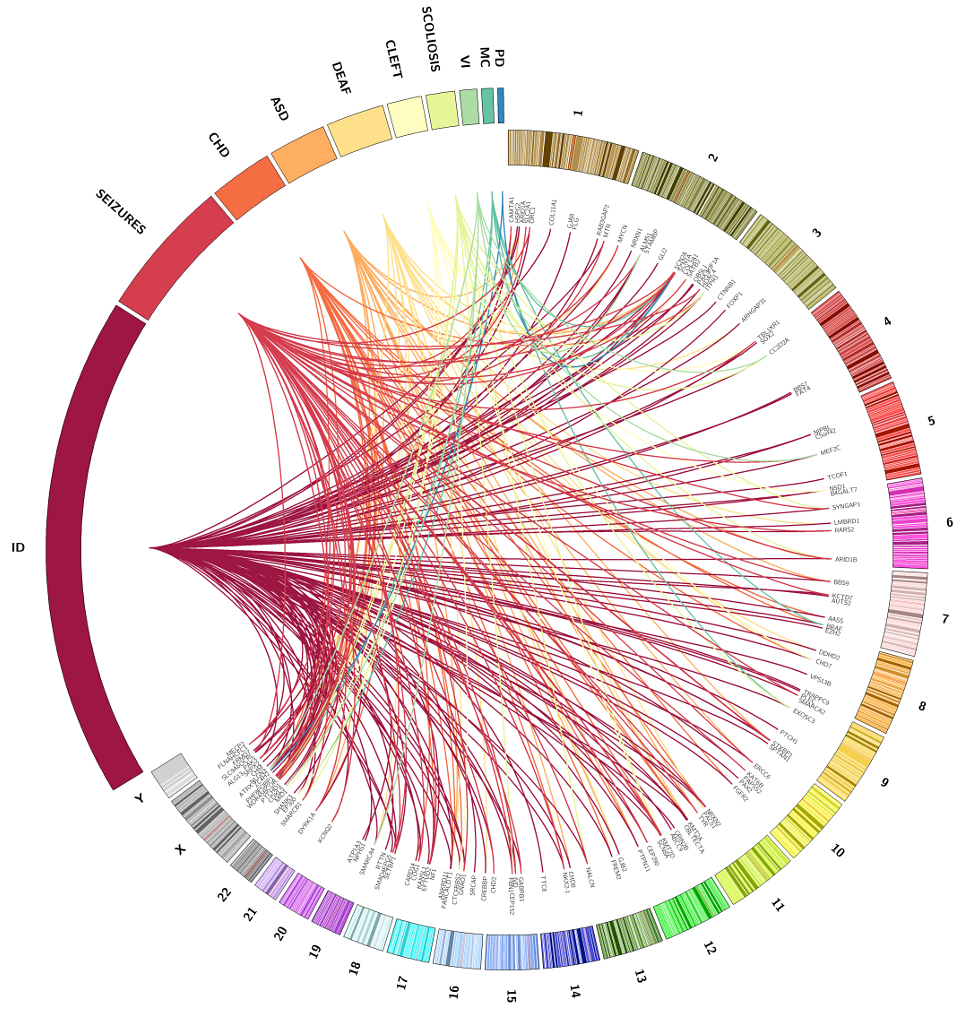
The first nationwide project to genetically diagnose rare diseases will pave the way for translating advances in genomics into patient care in the NHS. Deciphering Developmental Disorders (DDD), which is funded by the Wellcome Trust Sanger Institute and the Department of Health and Wellcome Trust through the Health Innovation Challenge Fund, is working with 12,000 families to diagnose their child’s developmental disorder, demonstrating the feasibility and value of introducing large-scale sequencing diagnostics into health care.
The project, which will continue into next year, has so far found a diagnosis for nearly a third of the first 1000 families analysed, where previous genetic tests were unable to find the cause. The diagnoses have focused on the 1100 genes that have previously been recognised as a cause of developmental disorders. This success rate will continue to improve as more disease-causing genes are discovered.
“Each of these patient families has been through a long diagnostic odyssey before taking part in the DDD project. For many, we are able to offer the diagnosis they have waited so long for. Sometimes this improves clinical management, but simply knowing the source of the problem can provide families with peace of mind.”
Dr Helen Firth An author from the Department of Clinical Genetics at Addenbrooke’s Hospital
Genome-wide microarray and whole exome sequencing is used to identify nearly 80,000 variants in each child, a handful of which potentially hold the genetic key to their disorder. Filtering out benign changes and unrelated findings accurately and efficiently is the major challenge facing researchers. A key step in this process is to compare symptoms identified by clinicians, such as neurodevelopmental delay or abnormal growth, with an in-house database of all known genes associated with developmental disorders.
“This project requires the collaborative efforts of hundreds of NHS staff and researchers. Clinicians managing patients on the frontline, database curators ploughing through the latest research, bioinformaticians developing computational tools and genetics experts interpreting results are all working together towards the common goal of finding answers for these patients.”
Dr David FitzPatrick an author from the Medical Research Council’s Human Genetics Unit in Edinburgh
Most of the causative genetic changes are new mutations in the affected child that were not inherited from either parent. Where unaffected parents were sequenced as well as the child Researchers saw a 10-fold reduction in the number of variants needing clinical evaluation. This study shows the importance of using parental genetic data, where possible, for the diagnosis of rare disorders.
The DDD study will continue to recruit families until April 2015, and will continue to try to find a genetic diagnosis for all the remaining undiagnosed families. It is hoped that the approach developed in this study will be rapidly incorporated in standard clinical practice to maximise the diagnosis of rare genetic disorders.
“By pulling together 24 regional genetics services and more than 180 clinical geneticists, we’ve implemented one single exome sequence diagnostic pipeline for children across the country. The project has shown that large-scale genome-wide testing, which brings such enormous benefits to patients and families, is both effective and affordable.”
Dr Caroline Wright, lead author and Programme Manager for the Deciphering Developmental Disorders project
The research is published in The Lancet.
More information
Funding
The DDD study presents independent research commissioned by the Health Innovation Challenge Fund [grant number HICF-1009-003], a parallel funding partnership between the Wellcome Trust and the Department of Health and the Wellcome Trust Sanger Institute [grant number WT098051].
Participating Centres
Wellcome Trust Sanger Institute, University of Oxford, University of Edinburgh, Cambridge University Hospitals Foundation Trust and all NHS regional genetics services in the UK and Republic of Ireland.
Publications:
Selected websites
Deciphering Developmental Disorders
The DDD study presents independent research commissioned by the Wellcome Trust Sanger Institute (grant ref WT098051) the Health Innovation Challenge Fund (grant ref HICF-1009-003), a parallel funding partnership between the Department of Health and Wellcome Trust.
Health Innovation Challenge Fund
The Health Innovation Challenge Fund is positioned as a translational funding scheme to accelerate the clinical application of projects that are well advanced along the development pathway. Its objectives are to:
- Stimulate the development and uptake of innovative products, technologies and interventions for the benefit of patients in the NHS and other healthcare systems.
- Support UK-led projects, which target unmet, or poorly met, healthcare needs.
- Provide translational funding for projects that have demonstrated ‘proof-of-principle’ and have the potential for early clinical use or adoption;
- Take the product, technology or intervention to the stage at which it is sufficiently developed to be attractive to follow-on funders or investors.
- Encourage the collaboration of companies, academia and clinicians to better confront today’s healthcare challenges.
The Department of Health
The Department of Health (DH) helps people to live better for longer. We lead, shape and fund health and care in England, making sure people have the support, care and treatment they need, with the compassion, respect and dignity they deserve. DH is a ministerial department, supported by 25 agencies and public bodies.
The Wellcome Trust Sanger Institute
The Wellcome Trust Sanger Institute is one of the world’s leading genome centres. Through its ability to conduct research at scale, it is able to engage in bold and long-term exploratory projects that are designed to influence and empower medical science globally. Institute research findings, generated through its own research programmes and through its leading role in international consortia, are being used to develop new diagnostics and treatments for human disease.
The Wellcome Trust
The Wellcome Trust is a global charitable foundation dedicated to achieving extraordinary improvements in human and animal health. We support the brightest minds in biomedical research and the medical humanities. Our breadth of support includes public engagement, education and the application of research to improve health. We are independent of both political and commercial interests.