Bugs without borders
Researchers show that the global epidemic of Clostridium difficile 027/NAP1/BI in the early to mid-2000s was caused by the spread of two different but highly related strains of the bacterium rather than one as was previously thought. The spread and persistence of both epidemics were driven by the acquisition of resistance to a frontline antibiotic.
Unlike many other healthcare-associated bacteria, C. difficile produces highly resistant and infectious spores. These spores can promote the transmission of C. difficile and potentially facilitates its spread over greater geographical distances, even across continents.
This study highlights the ease and rapidity with which the hospital bacterium, C. difficile, can spread throughout the world, emphasising the interconnectedness of the global healthcare system.
“Between 2002 and 2006, we saw highly publicised outbreaks of C. difficile in hospitals across the UK, USA, Canada and Europe. We used advanced DNA sequencing to determine the evolutionary history of this epidemic and the subsequent pattern of global spread.
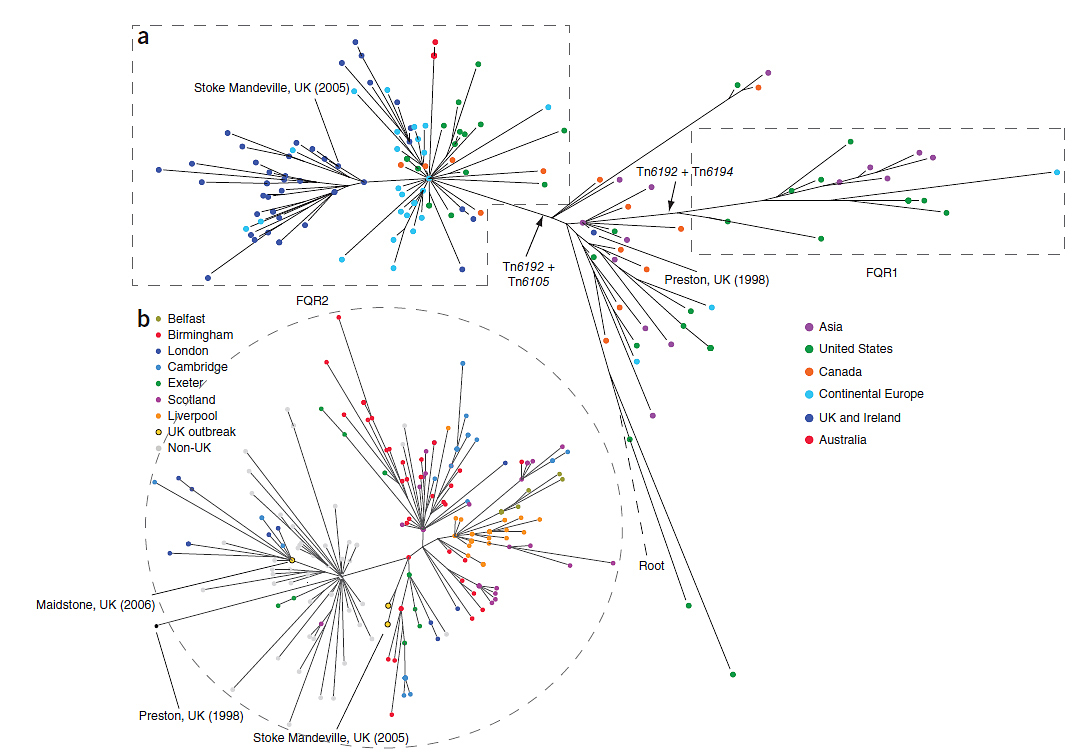
“We found that this outbreak came from two separate epidemic strains or lineages of C. difficile, FQR1 and FQR2, both emerging from North America over a very short period and rapidly spread between hospitals around the world.”
Dr Miao He First author from the Wellcome Trust Sanger Institute
The team used the genetic history to map both epidemic strains of C. difficile using a global collection of samples from hospital patients between 1985 and 2010. They demonstrated that the two C. difficile strains acquired resistance to this antibiotic, fluoroquinolone, separately, a key genetic change that may have instigated the epidemics in the early 2000s.
“Up until the early 2000s, fluoroquinolone was an effective treatment for C. difficile infection. We’ve seen that since these strains acquired resistance to this frontline antibiotic, not only is it now virtually useless against this organism, but resistance seems to have been a major factor in the continued evolution and persistence of these strains in hospitals and clinical settings.”
Professor Brendan Wren Author from the London School of Hygiene and Tropical Medicine
The team found the first outbreak strain of C. difficile, FQR1 originated in the USA and spread across the country. They also saw sporadic cases of this strain of C. difficile in Switzerland and South Korea. They found that the second strain of C. difficile, FQR2, originated in Canada and spread rapidly over a much wider area, spreading throughout North America, Australia and Europe.
The team showed that the spread of C. difficile into the UK was frequently caused by long-range geographical transmission event and then spread extensively within the UK. They confirmed separate transmission events to Exeter, Ayrshire and Birmingham from North America and a transmission event from continental Europe to Maidstone. These events triggered large-scale C. difficile outbreaks in many hospitals across the UK in the mid-2000s.
“We have exposed the ease and rapidity with which these fluoroquinolone-resistant C. difficile strains have transmitted across the world. Our research highlights how the global healthcare system is interconnected and how we all need to work together when an outbreak such as this occurs.
“Our study heralds a new era of forensic microbiology for the transmission tracking of this major global pathogen and will now help us understand at the genetic level how and why this pathogen has become so aggressive and transmissible worldwide. This research will act as a database for clinical researchers to track the genomic changes in C. difficile outbreaks.”
Dr Trevor Lawley Lead author from the Wellcome Trust Sanger Institute
More information
Funding
This project was funded by the Wellcome Trust, a Medical Research Council New Investigator Research Grant and the Scottish Infection Research Network.
Participating Centres
A full list of participating centres can be found in the paper.
Publications:
Selected websites
The Wellcome Trust Sanger Institute
The Wellcome Trust Sanger Institute is one of the world’s leading genome centres. Through its ability to conduct research at scale, it is able to engage in bold and long-term exploratory projects that are designed to influence and empower medical science globally. Institute research findings, generated through its own research programmes and through its leading role in international consortia, are being used to develop new diagnostics and treatments for human disease.
The Wellcome Trust
The Wellcome Trust is a global charitable foundation dedicated to achieving extraordinary improvements in human and animal health. We support the brightest minds in biomedical research and the medical humanities. Our breadth of support includes public engagement, education and the application of research to improve health. We are independent of both political and commercial interests.


