Nobody's perfect
Researchers at Cambridge and Cardiff have found that, on average, a normal healthy person carries approximately 400 potentially damaging DNA variants and two variants known to be associated directly with disease traits. They showed that one in ten people studied is expected to develop a genetic disease as a consequence of carrying these variants.
It has been known for decades that all people carry some damaging genetic variants that appear to cause little or no ill effect. But this is the first time that researchers have been able to quantify how many such variants each of us has, and list them. This figure of 400 is likely to increase as more and more powerful genetic studies discover rare genetic variants more efficiently. Such research brings to the fore ethical questions surrounding anonymous studies and incidental findings.
“For over half a century, medical geneticists have wanted to establish the magnitude of the damage caused by harmful variants in our genomes. Our study finally brings us closer to understanding the extent of these damaging mutations.
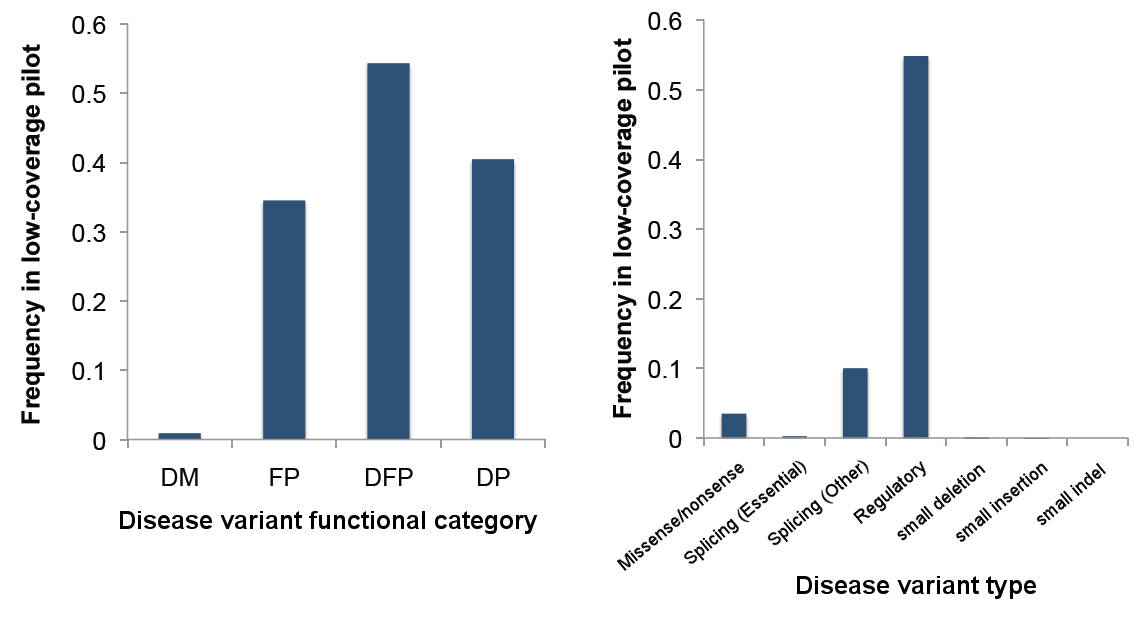
“We measured the number of potentially damaging variants in the genomes of apparently normal healthy humans by comparing two different datasets: whole genome sequences from 179 people in the 1000 Genomes Pilot Project, who were unlikely to have any overt genetic disease at the time of sampling, and information from the Human Gene Mutation Database (HGMD), a detailed catalogue of human disease-causing mutations that have been reported in the scientific literature.”
Dr Yali Xue Lead author from the Wellcome Trust Sanger Institute
In many cases, the disease or damaged variants were single, ‘recessive’ genetic variants that are unlikely to cause any harm to the carrier. A recessive genetic variant will only exert its effect when two copies – one in each chromosome – are present.
In one in ten people, however, the team could point to a potential clinical effect of the genetic variants. This is because these people either carry two copies of a specific recessive disease variant, or alternatively a dominant genetic variant. Dominant disease genetic variants can give rise to a disease trait when even a single copy is present.
“In the majority of people we found to have a potential disease-causing mutation, the genetic condition is actually quite mild, or would only become apparent in the later decades of life. We now know that normal healthy people can possess many damaged or even completely inactivated proteins without any noticeable impact on their health. It is extremely difficult to predict the clinical consequences of a given genetic variant, but databases such as HGMD promise to come into their own as we enter the new era of personalized medicine.”
Professor David Cooper Lead author of the study from Cardiff University
Catalogues of disease-causing variants such as HGMD have been created over the past two decades but they are still far from complete. Disease variants are generally extremely rare and comprehensive searches for such mutations in many populations have scarcely begun.
The genome samples selected for this study were anonymized so the participants could not receive any information about whether or not they might be at risk for a particular genetic disorder. This is increasingly becoming an ethical issue for medical geneticists.
“Should incidental findings be fed back to people who have volunteered their sample to a study? There is no clear answer to this question. All of our genomes contain flaws; some of us will carry deleterious variants but will not be at risk of acquiring the associated disease for one reason or another. For others, there will be health consequences, and early warning could be useful, but might still come as an unwelcome surprise to the participant.”
Dr Chris Tyler-Smith Lead author from the Wellcome Trust Sanger Institute
As DNA sequencing becomes more commonplace, geneticists must determine the most ethical way to handle this sensitive information.
More information
Funding
This study was funded by the Wellcome Trust and BIOBASE GmbH.
Participating Centres
- The Wellcome Trust Sanger Institute, Hinxton, Cambridge CB10 1SA, UK
- Institute of Medical Genetics, School of Medicine, Cardiff University, Heath Park, Cardiff CF14 4XN, UK
Publications:
Selected websites
The Wellcome Trust Sanger Institute
The Wellcome Trust Sanger Institute is one of the world’s leading genome centres. Through its ability to conduct research at scale, it is able to engage in bold and long-term exploratory projects that are designed to influence and empower medical science globally. Institute research findings, generated through its own research programmes and through its leading role in international consortia, are being used to develop new diagnostics and treatments for human disease.
The Wellcome Trust
The Wellcome Trust is a global charitable foundation dedicated to achieving extraordinary improvements in human and animal health. We support the brightest minds in biomedical research and the medical humanities. Our breadth of support includes public engagement, education and the application of research to improve health. We are independent of both political and commercial interests.