A leap forward for red blood cell formation
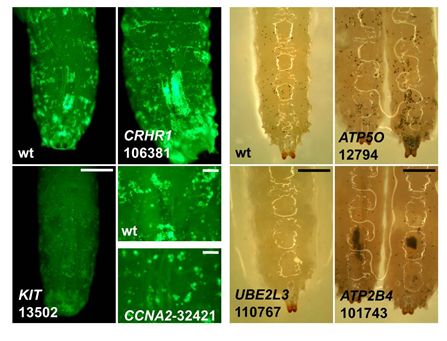
New research is revealing how red blood cells are made and how the body regulates the amount of haemoglobin that is packaged in red blood cells at any time. Genomic analysis techniques have doubled the number of genetic regions that are likely to be involved in red blood cell formation and subsequent study using fruit flies has given insights into what these regions do.
Haemoglobin is the protein which captures oxygen from the lungs for transport and delivery to tissues. It colours blood cells red and each day hundreds of millions of fresh red blood cells have to be formed by blood stem cells to replace the ones which come to the end of their life cycle. Anaemia, one of the most common disorders for which people visit their surgery, ensues if the production of new red blood cells is insufficient or their lifespan is shortened. The new genetic information is laying the foundations for future studies into the roots of anaemia by uncovering new biological pathways and mechanisms involved in controlling the size and number of red blood cells and the levels of haemoglobin.
The researchers used genome-wide association studies to identify genetic regions that appeared to influence the formation of red blood cells and their haemoglobin content.
“We studied the genetic influences behind six different physical parameters of red blood cells that reflect the volume and number of red blood cells and the levels of haemoglobin. Our initial genetic association study looked into the genomes of 135,367 people and identified 75 genetic regions that directly influence these different traits of red blood cells. More than half – 43 – of these discoveries are new in people.”
Dr John Chambers Lead author from Imperial College, London
The team then closely examined using computational biology approaches the 75 genetic regions and the more than 3,000 genes responsible for protein production lie close to these regions. They prioritised 121 ‘candidate’ genes or genes that are likely to regulate a trait in red blood cells from this list and investigated their function using information on model systems like from public databases as well as newly-generated data for fruit fly.
“Our work shows how model systems like fruit fly and mice can be used to provide insights into human genetics. We searched through a Mouse Genome database and found that 29 of our 121 candidate genes are linked to red blood cell formation in mice.
“These previous studies revealed that – when the function of these genes was switched off- the mice frequently developed reduced numbers of red blood cells, and anaemia. These observations made in mice make it highly likely that the remaining candidate genes, about which there is no knowledge yet, are also important regulators of red blood cell formation in people.”
Professor Willem Ouwehand Lead author from the University of Cambridge and NHS Blood and Transplant
To investigate further, the team then reduced or ‘silenced’ the activity of the candidate genes in fruit flies. Although fruit flies do not have red blood cells, they share some of the gene functions leading to the formation of blood elements. These studies confirmed that sets of genes involved in controlling human red blood cell traits in people were also important for the formation of blood cells in fly.
“These results support the view that genetic association studies identify sets of genes that are conserved in evolution across a wide range of species. This is exciting because it means that we can obtain extensive new insights into the genetics and biological pathway of human health by studying model organisms.
“Although the underlying mechanisms for the majority of genes we’ve identified still need to be elucidated, our research is opening many doors for future studies on the generation of red blood cells for clinical use in the laboratory and may also provide insights which may lead to improvements in the treatment of patients with inherited anaemias.”
Dr Nicole Soranzo Lead author from the Wellcome Trust Sanger Institute
Funding
A full list of funding can be found on the paper.
Participating Centres
A full list of participating centres can be found on the paper.
More information
Publications:
Selected websites
Imperial College London
Imperial College London consistently rated amongst the world’s best universities, Imperial College London is a science-based institution with a reputation for excellence in teaching and research that attracts 14,000 students and 6,000 staff of the highest international quality. Innovative research at the College explores the interface between science, medicine, engineering and business, delivering practical solutions that improve quality of life and the environment – underpinned by a dynamic enterprise culture.
Since its foundation in 1907, Imperial’s contributions to society have included the discovery of penicillin, the development of holography and the foundations of fibre optics. This commitment to the application of research for the benefit of all continues today, with current focuses including interdisciplinary collaborations to improve global health, tackle climate change, develop sustainable sources of energy and address security challenges.
In 2007, Imperial College London and Imperial College Healthcare NHS Trust formed the UK’s first Academic Health Science Centre. This unique partnership aims to improve the quality of life of patients and populations by taking new discoveries and translating them into new therapies as quickly as possible.
NHS Blood and Transplant
NHS Blood and Transplant (NHSBT) is a joint England and Wales Special Health Authority. Its remit includes the provision of a reliable, efficient supply of blood and associated services to the NHS in England and North Wales. It is also the organ donor organisation for the UK and is responsible for matching and allocating donated organs. Eight research themes make up NHSBT’s planned Research & Development Strategy. The main research sites, in collaboration with University partners, are Cambridge, Oxford, London and Bristol, with specialist activity in Birmingham and Liverpool.
University of Cambridge
The mission of the University of Cambridge is to contribute to society through the pursuit of education, learning and research at the highest international levels of excellence. It admits the very best and brightest students, regardless of background, and offers one of the UK’s most generous bursary schemes. The University of Cambridge’s reputation for excellence is known internationally and reflects the scholastic achievements of its academics and students, as well as the world-class original research carried out by its staff. Some of the most significant scientific breakthroughs occurred at the University, including the splitting of the atom, invention of the jet engine and the discoveries of stem cells, plate tectonics, pulsars and the structure of DNA. From Isaac Newton to Stephen Hawking, the University has nurtured some of history’s greatest minds and has produced more Nobel Prize winners than any other UK institution with over 80 laureates.
The Wellcome Trust Sanger Institute
The Wellcome Trust Sanger Institute is one of the world’s leading genome centres. Through its ability to conduct research at scale, it is able to engage in bold and long-term exploratory projects that are designed to influence and empower medical science globally. Institute research findings, generated through its own research programmes and through its leading role in international consortia, are being used to develop new diagnostics and treatments for human disease.
The Wellcome Trust
The Wellcome Trust is a global charitable foundation dedicated to achieving extraordinary improvements in human and animal health. We support the brightest minds in biomedical research and the medical humanities. Our breadth of support includes public engagement, education and the application of research to improve health. We are independent of both political and commercial interests.