The pig genome
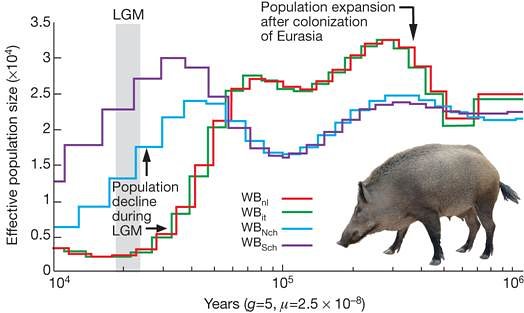
Their study found significant genetic differences between wild boar from Asia and Europe, which split from a common ancestor around a million years ago.
The mapping, sequencing and annotation teams at the Sanger Institute were key players in the international consortium to sequence and analyse the genome of the domestic pig. The results were published in Nature on Thursday 15 November.
The Swine Genome Sequencing Consortium supported the genome sequencing project, the bulk of which was carried out at the Wellcome Trust Sanger Institute. The project produced a high-quality draft sequence of a domesticated pig breed.
The scientists identified about 21,000 genes in the pig genome and compared these genes to their counterparts in human, mice, dogs, horses and cows to provide an understanding of both evolution and swine biology.
“The Sanger Institute mapping and cytogenetics, sequencing and annotation teams played a vital role in developing this important science. Our standards of sequence production have led to a high-quality genome from which our annotation team and collaborators have uncovered so much about the biology written in the pig genome.”
Carol Churcher Leader of the pig genome work at the Sanger Institute
The team discovered new details of common pig evolution after the ancestors of the domestic pig, which most resembled today’s wild boars, first emerged in Southeast Asia and gradually migrated across Eurasia.
“This understanding of the genetic origins of modern pigs is important as we breed pigs to meet growing demand more efficiently and to resist old and emerging diseases.”
Alan Archibald A professor of The Roslin Institute at the University of Edinburgh and a principal investigator on the study
Some gene families are undergoing relatively fast evolution in the domestic pig, with immune genes and olfactory genes quickly expanding. The pig has a greater number of unique olfactory genes than humans, mice or dogs, the researchers report.
And while pigs can smell a world of things humans and many other animals can’t, their sense of taste is somewhat impaired.
“The new analysis has important implications for agriculture, particularly since the domestic pig still has an ancestor-like wild cousin on the loose. Unlike the domestic cow, whose ancestors, the aurochs, are now extinct, the porcine lineage has a lot of genetic diversity remaining. We can easily find genes that might be still in the wild that we could use for breeding purposes today.”
Professor Lawrence Shook Of the University of Illinois
The new analysis also supports the use of the pig in studies of human diseases.
“In total, we found 112 positions where the porcine protein has the same amino acid that is implicated in a disease in humans,” the researchers wrote.
“This knowledge will be invaluable for pig breeding and exploring fundamental questions in biology and evolution. We identified many more gene variants implicated in human disease, further supporting the pig as a valuable biomedical model.”
Professor Martien Groenen Of Wageningen University
Some of the protein aberrations pigs share with humans are associated with obesity, diabetes, dyslexia, Parkinson’s disease and Alzheimer’s disease, the researchers report.
The study involved more than 40 institutions in 12 countries. It was funded by the United States Department of Agriculture, the European Commission, the Biotechnology and Biological Sciences Research Council, The Wellcome Trust as well as pig industry groups in Europe and the United States.
More information
Funding
A full list of supporting agencies can be found at the Nature website.
Participating Centres
A full list of participating centres can be found at the Nature website.
Publications:
Selected websites
The Wellcome Trust Sanger Institute
The Wellcome Trust Sanger Institute is one of the world’s leading genome centres. Through its ability to conduct research at scale, it is able to engage in bold and long-term exploratory projects that are designed to influence and empower medical science globally. Institute research findings, generated through its own research programmes and through its leading role in international consortia, are being used to develop new diagnostics and treatments for human disease.
The Wellcome Trust
The Wellcome Trust is a global charitable foundation dedicated to achieving extraordinary improvements in human and animal health. We support the brightest minds in biomedical research and the medical humanities. Our breadth of support includes public engagement, education and the application of research to improve health. We are independent of both political and commercial interests.


