When one is not enough: finding hyper-sensitive genes
Normally children inherit two versions of every gene: one from their mother and one from their father. This allows a gene to keep working even if one version is lost. However, more than 300 genes have been discovered that will cause developmental disorders if only one version of the gene is present – and more have yet to be identified. In this study, scientists compared these sensitive genes with more than 1000 non-sensitive genes to find common genetic ‘signposts’ that could then be used to predict whether a gene is likely to be sensitive or not.
“When we compared genes sensitive to the loss of single copy with non-sensitive genes, we found that there were key evolutionary and functional similarities between the sensitive genes. In particular, these genes tend to be longer and to be highly conserved among higher primates. Also we found that they tend to play a role in the early stages of embryonic development and often work in conjunction with other sensitive genes.”
Matt Hurles Leader of the study, from the Wellcome Trust Sanger Institute
The study’s authors looked at the genomes of more 8,000 apparently healthy people, underscoring the value of using a comprehensive understanding of normal variation to better identify disease-causing variation. They hope that these findings will enable researchers to quickly identify other sensitive genes and target them for further investigation.
“We have built a predictive computer model using the genetic signatures we identified and have rated more than half of all human genes with their probability of being sensitive to loss of one copy. Using this probability map, we expect that fellow researchers will be able to prioritise variants and genes for follow up studies.”
Ni Huang PhD student and first author on the publication, from the Wellcome Trust Sanger Institute
They also hope that this research will enable more accurate diagnosis of developmental disorders.
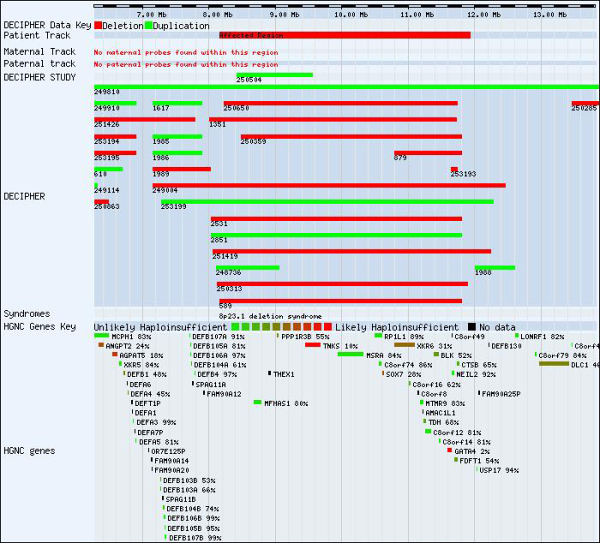
“Until now, clinicians have tried to diagnose developmental disorders by using the size of a chromosomal deletion, counting the number of genes lost and looking at the function of the genes where this is known. However, this does not take into account the underlying sensitivity of the deleted genes. Using our sensitivity predictions, we are better able to discriminate between deletions we know to cause childhood developmental disorders and those that do not.”
Helen Firth Consultant Clinical Geneticist, Addenbrooke’s Hospital, Cambridge
Every individual genome, not just in children with developmental disorders but even in healthy individuals, has more than a hundred genes that have lost one of their functioning copies. Understanding which genes are sensitive and which are not, allows us to interpret more fully personal genomes.
It is not currently well understood why certain genes require both versions to be working to maintain healthy function, but by identifying commonalities between them it is hoped that this will help shed more light on this mystery.
More information
Participating Centres
- The Wellcome Trust Sanger Institute, Wellcome Trust Genome Campus, Hinxton, Cambridge, United Kingdom
- Institute for Cellular and Molecular Biology, University of Texas, Austin, Texas, USA
- College of Life Science and Biotechnology, Yonsei University, Seoul, South Korea
Publications:
Selected websites
The Wellcome Trust Sanger Institute
The Wellcome Trust Sanger Institute, which receives the majority of its funding from the Wellcome Trust, was founded in 1992. The Institute is responsible for the completion of the sequence of approximately one-third of the human genome as well as genomes of model organisms and more than 90 pathogen genomes. In October 2006, new funding was awarded by the Wellcome Trust to exploit the wealth of genome data now available to answer important questions about health and disease.
The Wellcome Trust
The Wellcome Trust is a global charitable foundation dedicated to achieving extraordinary improvements in human and animal health. We support the brightest minds in biomedical research and the medical humanities. Our breadth of support includes public engagement, education and the application of research to improve health. We are independent of both political and commercial interests.


