Researchers Expand Efforts to Explore Functional Landscape of the Human Genome
Dr Tim Hubbard, Head of Informatics at the Sanger Institute, will lead efforts identify gene features across all 3 billion bases of DNA code in the human genome, the only scale-up programme awarded funding that is based outside the US.
“The pilot phase of ENCODE revealed a great deal, yet showed we still have much to learn about the content and the actions of the human genome. By expanding our efforts to the entire genome, we expect to discover many more genome features that will help us to understand human genome biology and the role of genome elements in health and disease.”
Dr Tim Hubbard Head of Informatics at the Sanger Institute
Dr Hubbard will lead an international team that will integrate a range of computational and informational studies of genes to develop accurate ‘annotation’ of the genome – the description of gene locations and structures. The project team will provide additional information about the strength of gene annotation and, by combining informatics and experimental approaches, will enhance the quality of catalogue of human genes.
“We have confidence that through this integrated project we will be able to eliminate many of the remaining uncertainties about the precise location of genes, their components and transcript structure in the human genome.”
Dr Tim Hubbard Head of Informatics at the Sanger Institute
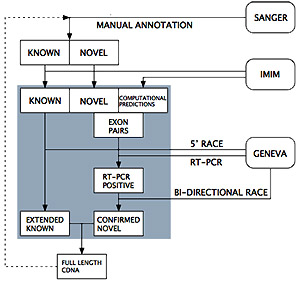
The project will build on the manual gene annotation work of the Sanger HAVANA group, led by Dr Jen Harrow, which played a major part in the ENCODE pilot and the automatically generated human gene annotation from the joint Sanger/EBI Ensembl project.
The new $80M ENCODE programme is a multidisciplinary programme that brings a range of cutting-edge research methods to the study of the genome, which were examined in the pilot phase. In addition to the research grants to support expansion of the ENCODE project, NHGRI also announced awards today for two pilot-scale projects, the establishment of an ENCODE data coordination centre and six projects to develop novel methods and technologies aimed at helping the ENCODE project achieve its goals.
“Based on ENCODE’s early success, we are moving forward with a full-scale initiative to build a parts list of biologically functional elements in the human genome. The ENCODE pilot produced findings that are reshaping many long-held views about our genome. ENCODE’s effort to survey the entire genome will uncover even more exciting surprises, providing us with a more complete picture of the biological roots of human health and disease.”
NHGRI Director Francis S. Collins MD, PhD
In June, the ENCODE research consortium published a set of landmark papers in the journals Nature and Genome Research that found the organization, function and evolution of the genome to be far more complicated than most had suspected. For example, while researchers have traditionally focused on studying genes and their associated proteins, the ENCODE data indicate the genome is a very complex, interwoven network in which genes are just one of many types of DNA sequences with functional impact.
“As was the case for the Human Genome Project and the ENCODE pilot, all of the data generated by the full-scale ENCODE project will be deposited into public databases as soon as they are experimentally verified. Free and rapid access to this data will enable researchers around the world to pose new questions and gain new insights into how the human genome functions.”
Peter Good PhD Programme director for genome informatics in NHGRI’s Division of Extramural Research
More information
ENCODE Scale-up Grantees (total $65.6M)
- Bradley Bernstein, Broad Institute of MIT and Harvard, Cambridge, MA, USA; High-Throughput Sequencing of Chromatin Regulatory Elements
- Gregory Crawford, Duke University Institute for Genome Sciences & Policy, Durham, NC, USA; Comprehensive Identification of Active Functional Elements in Human Chromatin
- Thomas Gingeras, Affymetrix Inc., Santa Clara, CA, USA; Comprehensive Characterization and Classification of the Human Transcriptome
- Tim Hubbard, Wellcome Trust Sanger Institute, Hinxton, UK; Integrated Human Genome Annotation: Generation of a Reference Gene Set
- Richard Myers, Stanford University, Stanford, CA, USA; Global Annotation of Regulatory Elements in the Human Genome
- Michael Snyder, Yale University, New Haven, CN, USA; Production Center for Global Mapping of Regulatory Elements
- John Stamatoyannopoulos, University of Washington, Seattle, WA, USA; A Comprehensive Catalog of Human DNase I Hypersensitive Sites
Other funding is awarded for data coordination, additional pilot projects and new technology development
Participating Centres and principle investigators in the Integrated Annotation Programme
- Tim Hubbard, Wellcome Trust Sanger Institute, Genome Campus, Hinxton, Cambridge CB10 1SA, UK
- Alexandre Reymond, The University of Lausanne, Switzerland
- Roderic Guigo, Centre de Regulació Genòmica (CRG), Barcelona, Catalonia, Spain
- David Haussler, University of California, Santa Cruz (UCSC) Santa Cruz, California, USA
- Michael Brent, Washington University, St Louis (WashU), St Louis, USA
- Manolis Kellis, Massachusetts Institute of Technology (MIT), Boston, USA
- Mark Gerstein, Yale University (Yale), New Haven, USA
- Alfonso Valencia, Spanish National Cancer Research Centre (CNIO), Madrid, Spain
Selected websites
NHGRI
NHGRI is one of 27 institutes and centres at NIH, an agency of the Department of Health and Human Services. NHGRI’s Division of Extramural Research supports grants for research and for training and career development at sites nationwide.
The National Institutes of Health
The National Institutes of Health – "The Nation’s Medical Research Agency" is a component of the U.S. Department of Health and Human Services. It is the primary federal agency for conducting and supporting basic, clinical and translational medical research, and it investigates the causes, treatments and cures for both common and rare diseases.
The Wellcome Trust Sanger Institute
The Wellcome Trust Sanger Institute, which receives the majority of its funding from the Wellcome Trust, was founded in 1992. The Institute is responsible for the completion of the sequence of approximately one-third of the human genome as well as genomes of model organisms and more than 90 pathogen genomes. In October 2006, new funding was awarded by the Wellcome Trust to exploit the wealth of genome data now available to answer important questions about health and disease.
The Wellcome Trust
The Wellcome Trust is the largest charity in the UK. It funds innovative biomedical research, in the UK and internationally, spending around £500 million each year to support the brightest scientists with the best ideas. The Wellcome Trust supports public debate about biomedical research and its impact on health and wellbeing.


