Hub genes
 Most human genetic diseases are complex, arising not from the action of a single gene, but from interactions between combinations of genes. However, these interactions are extremely difficult to identify in humans. Simple animals contain many of the same genes as humans and so, by understanding how genes interact in less complex animals, scientists can understand how human genes interact to cause disease. For the first time, researchers have systematically mapped these interactions in an animal and identify global properties of genetic interactions.
Most human genetic diseases are complex, arising not from the action of a single gene, but from interactions between combinations of genes. However, these interactions are extremely difficult to identify in humans. Simple animals contain many of the same genes as humans and so, by understanding how genes interact in less complex animals, scientists can understand how human genes interact to cause disease. For the first time, researchers have systematically mapped these interactions in an animal and identify global properties of genetic interactions.
On Sunday 16 July 2006, in the online version of Nature Genetics, a team from the Wellcome Trust Sanger Institute described the complete mapping of interactions between 65,000 pairs of genes and identified a novel class of genes that play a role in many unrelated processes and so may play a role in many different diseases.
This small group of genes – called ‘hub’ genes – was found in a study of interaction in the nematode worm C. elegans. The team show that hub genes are conserved in other organisms and suggest that their normal function is to act as genetic buffers, minimizing the effects of mutations in other genes. Further development of genetic interaction maps should help to identify other members of this key class of genes.
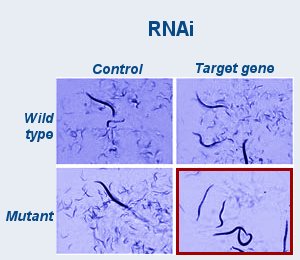
The researchers used RNA interference (RNAi), which is a method to suppress activity of individual genes. In the nematode, this can be done simply by feeding the worms bacteria containing the RNAi molecules. Worms can be quickly inspected for a more dramatic effect when two genes are suppressed, as compared with either gene alone.
“RNAi in the nematode is a particularly simple and elegant method to study the effects of genetic variation, and the worm is ideally suited to high-throughput analysis. If we want to understand how genes work in concert, we have to look at the activities of combinations of genes in living organisms.
“We found specific roles for many new genes in individual known pathways that are key in human disease but, more important, identified a class of hub genes, which affect many different pathways. We predict that other classes of hub genes will be identified using this method.”
Dr Andy Fraser Principal Investigator at the Wellcome Trust Sanger Institute
The team looked at more than 1700 genes known to be involved in three key cell functions communication between cells, control of gene activity and regulation of DNA-protein interaction. These activities are shared among animals and genes of these types are often mutated in human disease.
Normally biologists study gene function by mutating one gene at a time and looking at the consequences for the animal. In this study, researchers looked at the additional effects resulting from inactivating two genes simultaneously. To do this the researchers used RNAi to inactivate each of 1700 genes in worms that already carried a mutation in another gene. In total 37 different mutant worm strains were used, meaning that a total of 65,000 possible pairwise combinations were tested. Of these, 349 combinations gave an observable phenotype, defining genetic interactions between these genes. This set of interactions, derived from study of a restricted subset of genes, suggests that, on a genome-wide scale, there might be as many as 1 million pairwise combinations of mutations that could be lethal to the worm.
Although the majority of genes interacted with only 1-5 other genes, six genes had much higher numbers of interactions. All of these were involved in modifying the structure of chromatin, the malleable combination of DNA and protein that can serve to alter genetic activity.
“Our results suggest that genetic disease in humans, like the effects we see in the worm, could be due to the combined effect of two mutations, one in a gene specific to a single pathway, the other in one of these generic, widely acting hub genes. This way of looking at how combinations of mutations add to give clinical disease is very simple and is a great example of the kind of insights into human disease that can come from studies in more primitive animals like the worm.
“We found that hub genes could influence the effects of mutation in multiple, unrelated biochemical pathways. We are sure that our search has uncovered only a subset of these and our current research is designed to detect more.”
Dr Ben Lehner first author on the paper
The hub genes are not unique to the worm – all are found in other animals including human, suggesting that they play a fundamental role in modulating gene activity. Previous research has suggested that a gene called hsp90 acts as a buffer, or biochemical capacitor, to minimize naturally occurring genetic variation. Extreme effects of variation in activity of one gene might be harmful and limiting variation could play a vital role in survival of an organism.
Selected websites
The Wellcome Trust Sanger Institute
The Wellcome Trust Sanger Institute, which receives the majority of its funding from the Wellcome Trust, was founded in 1992. The Institute is responsible for the completion of the sequence of approximately one-third of the human genome as well as genomes of model organisms and more than 90 pathogen genomes. In October 2006, new funding was awarded by the Wellcome Trust to exploit the wealth of genome data now available to answer important questions about health and disease.
The Wellcome Trust and Its Founder
The Wellcome Trust is the most diverse biomedical research charity in the world, spending about £450 million every year both in the UK and internationally to support and promote research that will improve the health of humans and animals. The Trust was established under the will of Sir Henry Wellcome, and is funded from a private endowment, which is managed with long-term stability and growth in mind.


