One-tenth of our genome in one day: from sequence to disease to evolution
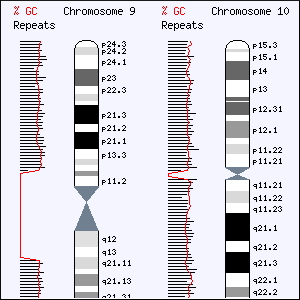
In two landmark reports published today in Nature (27th May 2004), researchers from Wellcome Trust Sanger Institute and their collaborators announce the findings of their study of the finished sequence of human chromosomes 9 and 10. Together, these chromosomes comprise one-tenth of our human genome.
The detailed analyses provide an overview of the chromosome landscape but, more important, identify new genes involved in disease and give a glimpse of our relationship to our nearest living relative, the chimpanzee. Biomedicine is already reaping benefit from the freely available sequences.
Chromosome 9
The finished sequence of chromosome 9 comprises nearly 110 million letters of DNA code (base-pairs). Using this sequence, the team have identified almost 1200 genes. Chromosome 9 harbours four genes that can cause sex-reversal, all the human interferon type 1 genes (interferon is important in suppressing cancer development and in resisting virus infection), a gene implicated in neurodegenerative disease (CHAC), as well as a gene (abl) that is involved in 90% of chronic myeloid leukaemia (CML) cases. CML is one of the first cancers for which a treatment has been developed based on understanding the human genome sequence.
Through remarkable efforts, the chromosome 9 team produced finished sequence from many duplicated regions that are normally refractile to mapping and sequencing.
“The overview of Chromosome 9 presents an unusual structural landscape: there are large regions of repeated sequence bearing evidence of duplications in our evolutionary history. In our detailed view we were able to identify a number of recently duplicated genes including those important in immune response.
“More important, genomic studies afford the first opportunity to see the whole picture. With expertly analysed chromosome sequences accessible to the scientific community, biomedicine can do research into common disease that simply was not possible before we reached this stage.”
Sean Humphray Leader of the Chromosome 9 team
“Our original interest in this chromosome grew out of our identification of several families around the world with an inherited predisposition to a specific bone cancer. We successfully linked the trait to chromosome 9p21-22 and now the completed gene sequence and annotations are leading us to identifying a novel sarcoma gene.
“Undoubtedly, the sequence will provide the basis for identification, characterization, and exploration of novel genes and the mechanisms by which normal and cancerous cells generate new and unexpected proteins from what were previously thought to be ‘well-characterized’ genes.”
Professor John A. Martignetti MD PhD Of the Departments of Human Genetics and Pediatrics, Mount Sinai School of Medicine
Chromosome 10
The team decoded 131 million base-pairs of the chromosome 10 sequence and found 1357 genes. Alterations in 85 of these genes are known to predispose to diseases such as a form of epilepsy (LGI1), obesity (GAD2), and cancer. The finished sequence is an essential tool in both analysing these genes and finding additional ones; there is genetic evidence for type I diabetes, schizophrenia and Alzheimer’s disease.
Comparisons with chimpanzee sequence revealed a surprisingly high degree of difference in the sequence of many genes.
“Our study painted one more section of the genomic landscape in humans but the sequence has still more tales to tell. Comparisons with genomes of other species begin to fill the evolutionary jigsaw and promise to give new clues – for example, the chimp data opens a window on the genome biology of primates. But an end point in our research is to understand disease. With an annotated high-quality sequence in hand, studies of sequence differences between individuals can begin to pick apart the foundation of many of the common but complex diseases.”
Panos Deloukas Leader of the Chromosome 10 team
Chromosome 10 offers a remarkable example as to how duplications have both shaped present day genomes and impacted on gene count. Also of interest is the number of genes sharing the same space; 15% of genes on chromosome 10 are found as overlapping pairs: conventionally it is thought that each gene occupies a unique region of DNA sequence.
Comparisons
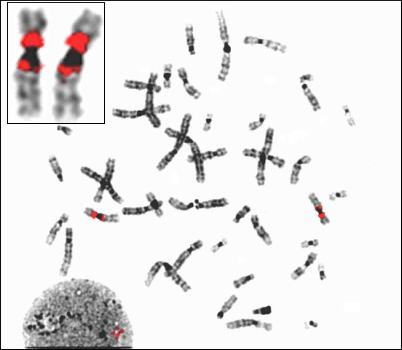
The two chromosome sequences point to the rich variety in our genome: chromosome 9 is peppered with regions copied from elsewhere in the genome: chromosome 10 is rich in overlapping genes.
When the teams compared human with all available chimpanzee sequence, a large number of differences was found – for chromosome 10, nearly half the differences would alter the protein sequence. Moreover, nearly 2% of differences would lead to a truncated (and possibly non-functional) protein. Among the changes that appear to be significant are those affecting a gene that plays a role in cognition and behaviour (HTR7) and one involved in development of the embryo (NODAL).
On chromosome 9, genes such as IL11RA, a gene involved in the immune response, and C9orf37 (a gene identified using the newly available finished sequence) showed a significant difference at the sequence level.
With publications of completed analysis of all chromosomes in the next year, our picture of genetic differences between the two species will become clearer.
“We have continued to improve the quality of our research and the resources we provide to the research community worldwide. These are hugely exciting times in biomedicine as we build on superb results, such as these chromosome studies, with new programmes to tackle human disease in ways that were not possible only ten years ago.
“Our quest to understand human health and disease will be long, and we must not raise expectations, but we can see now the directions we have to go. And because of genome sequence, we are seeing some of those possibilities for the first time.”
Professor Allan Bradley Director of The Wellcome Trust Sanger Institute
The Wellcome Trust Sanger Institute is responsible for the largest single contribution to the finished human genome sequence. It also, together with the European Bioinformatics Institute, provides the Ensembl web tool (http://www.ensembl.org/) which allows researchers free and unfettered access to genome sequence, variation information and annotation. Ensembl is a success story of the genomic age, receiving more than 3,000,000 hits each week.
More information
The analyses of the two sequences are published in Nature magazine on Thursday 27 May 2004. The papers are entitled: “DNA sequence and analysis of human chromosome 9” by Sean Humphray et al. and “The DNA sequence and comparative analysis of human chromosome 10” by Panos Deloukas et al.
Chromosome 9 Participating Centres
- The Wellcome Trust Sanger Institute, Wellcome Trust Genome Campus, Hinxton, Cambridge CB10 1SA, UK
- European Bioinformatics Institute, Wellcome Trust Genome Campus, Hinxton, Cambridge CB10 1SD, UK
- The Institute of Human Genetics, The International Centre for Life, University of Newcastle upon Tyne, Newcastle upon Tyne NE1 3BZ, UK
- German Research Centre for Biotechnology (GBF), Department of Genome Analysis, Mascheroder Weg 1, D-38124 Braunschweig, Germany
- HUGO Gene Nomenclature Committee, Department of Biology, University College London, Wolfson House, 4 Stephenson Way, London NW1 2HE, UK
Chromosome 10 Participating Centres
- The Wellcome Trust Sanger Institute, Wellcome Trust Genome Campus, Hinxton CB10 1SA, UK
- Genome Therapeutics Corporation, 100 Beaver Street, Waltham, Massachusetts 02453, USA
- Agencourt Bioscience Corporation, 100 Cummings Center, Beverly, Massachusetts 01915, USA
- Department of Biology, University of Crete & Institute of Molecular Biology and Biotechnology, Foundation of Research and Technology, PO Box 2208, 71409 Heraklion, Crete, Greece
- European Bioinformatics Institute (EBI), Wellcome Trust Genome Campus, Hinxton CB10 1SD, UK
- HUGO Gene Nomenclature Committee, Department of Biology, University College London, London NW1 2HE, UK
- Institute of Human Genetics, University Hospital Schleswig-Holstein Campus Kiel, Schwanenweg 24, D-24105 Kiel, Germany
Websites
Publications:
Selected websites
The Wellcome Trust and Its Founder
The Wellcome Trust is the most diverse biomedical research charity in the world, spending about £450 million every year both in the UK and internationally to support and promote research that will improve the health of humans and animals. The Trust was established under the will of Sir Henry Wellcome, and is funded from a private endowment, which is managed with long-term stability and growth in mind.
The Wellcome Trust Sanger Institute
The Wellcome Trust Sanger Institute, which receives the majority of its funding from the Wellcome Trust, was founded in 1992. The Institute is responsible for the completion of the sequence of approximately one-third of the human genome as well as genomes of model organisms and more than 90 pathogen genomes. In October 2006, new funding was awarded by the Wellcome Trust to exploit the wealth of genome data now available to answer important questions about health and disease.