COSMIC
COSMIC, the "Catalogue Of Somatic Mutations In Cancer" is an expert-curated database encompassing the wide variety of somatic mutation mechanisms causing human cancer.
About
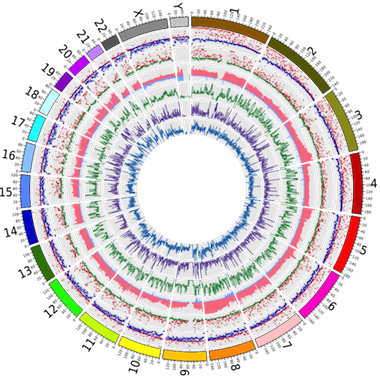 Hand-curation of key cancer genes (selected from the Cancer Gene Census) provide in-depth detail on mutation distributions and effects, whilst semi-automated curation of cancer genomes provides broad somatic annotations toward target discovery and identification of patterns and signatures. This information is fully available via website or download, updated every three months.
Hand-curation of key cancer genes (selected from the Cancer Gene Census) provide in-depth detail on mutation distributions and effects, whilst semi-automated curation of cancer genomes provides broad somatic annotations toward target discovery and identification of patterns and signatures. This information is fully available via website or download, updated every three months.
Curation
Cancer Gene Census: This is a list of hundreds of genes with substantial published evidence in oncology. Necessarily conservative, this is a very high-confidence list based on good-quality publications. Selection of high-impact genes from this list for curation drives COSMIC.
COSMIC: Upon selection of a gene from the Census for full expert curation, all papers mentioning its mutation in human cancer are collected and exhaustively curated before it is released into a new version of COSMIC. Once this initial curation is released, the gene is updated as significant new information is published. Each curator is responsible for a defined set of 60 or more genes, developing substantial expertise. In parallel, cancer genomes are curated via a more bioinformatic approach. Genomic data is obtained in standard formats; roughly half is from published supplementary information tables, and the other half from genome consortia such as TCGA, ICGC. Standard pipelines (eg Ensembl VEP) annotate these genomic data in genic terms for COSMIC release. Such molecular profiling includes point mutations, gene fusions, copy number annotations, structural breakpoints, gene expression and CpG island methylation variants.
Cancer Cell Lines Project: The Cell lines Project in COSMIC is an effort to fully profile over 1000 cell lines regularly used in cancer research; annotations include exome sequencing, CNV and gene expression profiling, RNASeq and CpG methylation. This information is maintained in a separate, but parallel system alongside COSMIC and regularly updated to highlight the most valuable information across the cell line panel.
Downloads
The COSMIC group make this information available in many ways to suit a variety of scientific and informatic users. Most accessibly, it can be viewed and analysed in its custom website (https://cancer.sanger.ac.uk), where distributions across genes, genomes and diseases can be explored easily and in detail. A full genome browser provides even greater information, correlating COSMIC mutations with a range of genomic annotations including promoters, ncRNAs and polymorphisms. Full downloads are also provided.
Further information
COSMIC is very intuitive and easy to use, we have provided substantial help pages explaining the various components and entry point into COSMIC. To further ease the navigation of the system, we also have Youtube Help tutorial videos.
COSMIC is now under licence, with academic access available freely and commercial access available for a licence fee. These funds are being used to expand the COSMIC group, particularly our curation team. As we enhance the number of genes under curation, and the scope of data we aim to capture, an expansion of this team will ensure that COSMIC continues to succeed in supporting a wide range of oncology research and product development.
Contact
If you need help or have any queries, please contact us using the details below.
Interested in receiving COSMIC news and release information? Register for a COSMIC account.
Please send all comments and suggestions to the cosmic@sanger.ac.uk
For licensing enquiries please contact cosmic-translation@sanger.ac.uk