The Weapons in Malaria's Evolutionary Arms Race
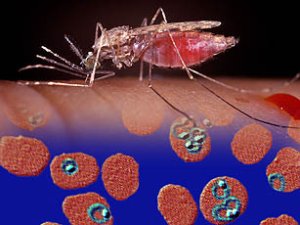 The numbers associated with malaria are awe-inspiring: 2.6 billion of the world’s people live in areas threatened by malaria; half a billion people are infected with the disease; more than one million people die each year from it.
The numbers associated with malaria are awe-inspiring: 2.6 billion of the world’s people live in areas threatened by malaria; half a billion people are infected with the disease; more than one million people die each year from it.
Today, a report led by the Wellcome Trust Sanger Institute details the weapons deployed by the parasite in this evolutionary arms race, and is published alongside two others in Nature Genetics. For the first time, we see how the genome of a clinical isolate is being shaped by the forces of our measures to battle malaria – a view that will reveal new areas to search for vaccine or drug targets.
There are many reasons – often related to poverty – why the burden of malaria has not been reduced. Simple solutions, such as provision of bednets, are required alongside a desperate need for new drugs to combat the parasite, Plasmodium falciparum, that causes malaria.
But this organism holds many, adaptable tools to help it to infect us and to evade our natural defence mechanisms and the treatments we develop to beat it.
The malaria parasite has many advantages in its battle to infect humans. Most significantly, it has the potential to ‘cloak’ itself in any one of a suite of cellular disguises though changes to its surface proteins. It also often lives as a mix of parasites in infected people, so that if we successfully treat one variant, another might rise from the background.
“We need new ways to look at the variety of Plasmodium surface proteins and its complex population structure. And these novel maps are new tools in the fight against malaria.
“For the first time, we have comprehensive maps that detail areas of variation – the regions where the effects of our defences are marked in the parasite genome. When we attack, the parasite responds and that is marked by the changes in the parasite DNA that we observe.”
Dr Matthew Berriman Co-leader of the project at Wellcome Trust Sanger Institute
 The report led by the Sanger Institute is joined by two others published online today (Sunday 10 December, 2006) in Nature Genetics – together they describe how the genome of the malaria parasite has been shaped by evolution and identify new areas to search for weaknesses in the parasite’s defences.
The report led by the Sanger Institute is joined by two others published online today (Sunday 10 December, 2006) in Nature Genetics – together they describe how the genome of the malaria parasite has been shaped by evolution and identify new areas to search for weaknesses in the parasite’s defences.
The Wellcome Trust Sanger Institute team produced the first complete sequence of a clinical isolate of Plasmodium: previous sequencing was been carried out on laboratory strains. They also produced partial sequence from a second isolate and from a strain that causes infection in chimpanzees. The teams led by Harvard School of Public Health and the Broad Institute of MIT and Harvard examined a set of lab strains, while the team led by the US National Institute of Allergy and Infectious Diseases examined four lab strains.
Previous work, including the sequence of the lab strain of Plasmodium falciparum identified regions of the genome, such as the var genes, that were variable within a strain. The new studies mapped more than 27,000 variants across the genome, showing other regions of the genome have a vital role to play.
In humans, Plasmodium lives in liver cells and red blood cells and it is this latter stage that determines the clinical course of the disease.
“We found that genes that are active in red blood cells and genes that are predicted to interact with host cells were especially variable. The conclusion is that this variation is driven by the arms race between the parasite and our host defences. For Plasmodium, to adapt is to survive – to cause disease.
“Looking at the evolution of categories of genes can point to those that are essential to the parasite’s lifestyle and are also likely to be stable – and hence most valuable for new treatments.”
Daniel Jeffares Investigator at Wellcome Trust Sanger Institute and first author of the study
Many of the most variable genes have not been assigned a function: however, it is likely that many will be surface proteins, part of the Plasmodium cloak. A newly identified gene called PF10_0355 was the second-most variable: it shares many properties with genes that have been targeted as possible vaccine candidates.
The genome study has identified most genes that are likely to be subject to attack by our immune system and hence rapidly evolving. Good targets for long-lasting new treatments are likely to be found among the more constant genes of the parasite’s armoury.
“The parasite genome is very plastic and carries the scars of its battle against its three main challenges – our rapidly evolving human immune system, the defensive responses of the mosquito and the insecticides and drugs we use to challenge it.
“Our variation studies bring biology to the parasite genome, uncovering the secrets of Plasmodium without – but as a prelude to – work in the lab. Our overview of evolution points to those gene variants that are responsible for disease effects – some were expected, but some are surprises.”
Dr Manolis Dermitzakis Co-leader of the project from the Wellcome Trust Sanger Institute
Humans infected with malaria often carry several variant strains. A consequence is that vaccines to combat malaria are very difficult to develop and may become ineffective if the parasite switches its coat. Moreover, the range of parasite diversity means that resistance to new drugs can become rapidly established from small numbers of existing resistant organisms. Widespread resistance to chloroquine has developed in only 50 years.
The Plasmodium map of variation can be used alongside maps of human variation, such as the HapMap or that of copy number variation published in Nature recently by a team from the Sanger Institute, to understand how the genome of each has been moulded by the activities of the other.
“The human genome carries imprints of our history of infection by malaria. Similarly the genome of the malaria parasite shows how it interacts with the human immune system. Understanding these interactions is key to the development of effective vaccines against malaria.”
Dr Mark Walport Director of the Wellcome Trust
The new map was developed with biological expertise from researchers at St George’s, University of London and The Weatherall Institute of Molecular Medicine, University of Oxford, John Radcliffe Hospital, Oxford. It is a snapshot of Plasmodium evolution, and provides a wealth of information for the malaria community, for example, by identifying genes that evolve too rapidly to be good drug targets. It shows researchers where to search for new treatments and where to avoid.
And new treatments and new measures are needed if the global community is to help reduce the burden of this disease.
If malaria occurred in the UK at global rates, 24 million people would be at risk, almost 4 million people would be infected and 10,000-20,000 people, almost all children, would die each year. As a nation, we would be horrified and huge efforts would be made to combat the disease.
The reason for the large range of deaths is perhaps the biggest indictment – in the real world of malaria we just don’t know how many people die.
More information
Participating Centres
- Informatics Division and Pathogen Sequencing Unit, Wellcome Trust Sanger Institute, Wellcome Trust Genome Campus CB10 1SA, Hinxton, UK
- Biomedical Primate Research Centre, Lange Kleiweg 139, RIJSWIJK, Postbus 3306, 2280 GH RIJSWIJK, The Netherlands
- Department of Molecular Biology and Genetics, Cornell University, Ithaca, NY14853, USA
- St George’s, University of London, Cranmer Terrace, London SW17 ORE UK
- The Weatherall Institute of Molecular Medicine, University of Oxford, John Radcliffe Hospital, Oxford OX3 9DS, UK
Acknowledgements
- This study was funded by the Wellcome Trust through their support of the Pathogen Sequencing Unit and Dr Dermitzakis’ group at the Wellcome Trust Sanger Institute
- Wellcome Trust Sanger Institute Pathogen Sequencing teams generated the sequence data used in this study
Publications:
Selected websites
The Wellcome Trust Sanger Institute
The Wellcome Trust Sanger Institute, which receives the majority of its funding from the Wellcome Trust, was founded in 1992. The Institute is responsible for the completion of the sequence of approximately one-third of the human genome as well as genomes of model organisms and more than 90 pathogen genomes. In October 2006, new funding was awarded by
The Wellcome Trust and Its Founder
The Wellcome Trust is the most diverse biomedical research charity in the world, spending about £450 million every year both in the UK and internationally to support and promote research that will improve the health of humans and animals. The Trust was established under the will of Sir Henry Wellcome, and is funded from a private endowment, which is managed with long-term stability and growth in mind.


